Genomic Profile in a Non-Seminoma Testicular Germ-Cell Tumor Cohort Reveals a Potential Biomarker of Sensitivity to Platinum-Based Therapy
- PMID: 35565196
- PMCID: PMC9101377
- DOI: 10.3390/cancers14092065
Genomic Profile in a Non-Seminoma Testicular Germ-Cell Tumor Cohort Reveals a Potential Biomarker of Sensitivity to Platinum-Based Therapy
Abstract
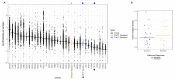
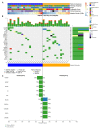
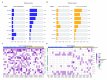
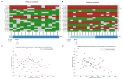
Despite having a favorable response to platinum-based chemotherapies, ~15% of Testicular Germ-Cell Tumor (TGCT) patients are platinum-resistant. Mortality rates among Latin American countries have remained constant over time, which makes the study of this population of particular interest. To gain insight into this phenomenon, we conducted whole-exome sequencing, microarray-based comparative genomic hybridization, and copy number analysis of 32 tumors from a Mexican cohort, of which 18 were platinum-sensitive and 14 were platinum-resistant. We incorporated analyses of mutational burden, driver mutations, and SNV and CNV signatures. DNA breakpoints in genes were also investigated and might represent an interesting research opportunity. We observed that sensitivity to chemotherapy does not seem to be explained by any of the mutations detected. Instead, we uncovered CNVs, particularly amplifications on segment 2q11.1 as a novel variant with chemosensitivity biomarker potential. Our data shed light into understanding platinum resistance in a Latin-origin population.
Keywords: CGH; CNVs and DNA breakpoints; SNVs; TGCT; WES; platinum-resistance.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

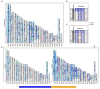
References
Grants and funding
LinkOut - more resources
Full Text Sources

