RB1-Negative Retinal Organoids Display Proliferation of Cone Photoreceptors and Loss of Retinal Differentiation
- PMID: 35565295
- PMCID: PMC9105736
- DOI: 10.3390/cancers14092166
RB1-Negative Retinal Organoids Display Proliferation of Cone Photoreceptors and Loss of Retinal Differentiation
Abstract
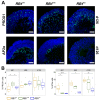
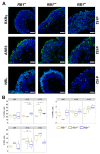
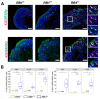
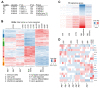
Retinoblastoma is a tumor of the eye in children under the age of five caused by biallelic inactivation of the RB1 tumor suppressor gene in maturing retinal cells. Cancer models are essential for understanding tumor development and in preclinical research. Because of the complex organization of the human retina, such models were challenging to develop for retinoblastoma. Here, we present an organoid model based on differentiation of human embryonic stem cells into neural retina after inactivation of RB1 by CRISPR/Cas9 mutagenesis. Wildtype and RB1 heterozygous mutant retinal organoids were indistinguishable with respect to morphology, temporal development of retinal cell types and global mRNA expression. However, loss of pRB resulted in spatially disorganized organoids and aberrant differentiation, indicated by disintegration of organoids beyond day 130 of differentiation and depletion of most retinal cell types. Only cone photoreceptors were abundant and continued to proliferate, supporting these as candidate cells-of-origin for retinoblastoma. Transcriptome analysis of RB1 knockout organoids and primary retinoblastoma revealed gain of a retinoblastoma expression signature in the organoids, characterized by upregulation of RBL1 (p107), MDM2, DEK, SYK and HELLS. In addition, genes related to immune response and extracellular matrix were specifically upregulated in RB1-negative organoids. In vitro retinal organoids therefore display some features associated with retinoblastoma and, so far, represent the only valid human cancer model for the development of this disease.
Keywords: RNA-seq; retinal organoids; retinoblastoma; stem cells.
Conflict of interest statement
The authors declare no conflict of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript, or in the decision to publish the results.
Figures



References
-
- Kooi I.E., Mol B.M., Massink M.P., Ameziane N., Meijers-Heijboer H., Dommering C.J., van Mil S.E., de Vries Y., van der Hout A.H., Kaspers G.J., et al. Somatic genomic alterations in retinoblastoma beyond RB1 are rare and limited to copy number changes. Sci. Rep. 2016;6:25264. doi: 10.1038/srep25264. - DOI - PMC - PubMed
-
- Rushlow D.E., Mol B.M., Kennett J.Y., Yee S., Pajovic S., Thériault B.L., Prigoda-Lee N.L., Spencer C., Dimaras H., Corson T.W., et al. Characterisation of retinoblastomas without RB1 mutations: Genomic, gene expression, and clinical studies. Lancet Oncol. 2013;14:327–334. doi: 10.1016/S1470-2045(13)70045-7. - DOI - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

