Development of 1-(4-(Substituted)piperazin-1-yl)-2-((2-((4-methoxybenzyl)thio)pyrimidin-4-yl)oxy)ethanones That Target Poly (ADP-Ribose) Polymerase in Human Breast Cancer Cells
- PMID: 35566199
- PMCID: PMC9100275
- DOI: 10.3390/molecules27092848
Development of 1-(4-(Substituted)piperazin-1-yl)-2-((2-((4-methoxybenzyl)thio)pyrimidin-4-yl)oxy)ethanones That Target Poly (ADP-Ribose) Polymerase in Human Breast Cancer Cells
Abstract
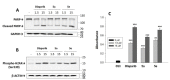
A number of uracil amides cleave poly (ADP-ribose) polymerase and therefore novel thiouracil amide compounds were synthesized and screened for the loss of cell viability in a human-estrogen-receptor-positive breast cancer cell line. The synthesized compounds exhibited moderate to significant efficacy against human breast cancer cells, where the compound 5e IC50 value was found to be 18 μM. Thouracil amide compounds 5a and 5e inhibited the catalytical activity of PARP1, enhanced cleavage of PARP1, enhanced phosphorylation of H2AX, and increased CASPASE 3/7 activity. Finally, in silico analysis demonstrated that compound 5e interacted with PARP1. Hence, specific thiouracil amides may serve as new drug-seeds for the development of PARP inhibitors for use in oncology.
Keywords: PARPi; apoptosis; breast cancer; thiouracil amides.
Conflict of interest statement
The authors declare no conflict of interest.
Figures











Similar articles
-
Design and Activity of Novel Oxadiazole Based Compounds That Target Poly(ADP-ribose) Polymerase.Molecules. 2022 Jan 21;27(3):703. doi: 10.3390/molecules27030703. Molecules. 2022. PMID: 35163965 Free PMC article.
-
Poly (ADP-ribose) polymerases inhibitor, Zj6413, as a potential therapeutic agent against breast cancer.Biochem Pharmacol. 2016 May 1;107:29-40. doi: 10.1016/j.bcp.2016.02.015. Epub 2016 Feb 24. Biochem Pharmacol. 2016. PMID: 26920250
-
Poly(ADP-ribose)polymerase (PARP) inhibition and anticancer activity of simmiparib, a new inhibitor undergoing clinical trials.Cancer Lett. 2017 Feb 1;386:47-56. doi: 10.1016/j.canlet.2016.11.010. Epub 2016 Nov 12. Cancer Lett. 2017. PMID: 27847302
-
Poly(ADP-ribose) Polymerase (PARP) and PARP Inhibitors: Mechanisms of Action and Role in Cardiovascular Disorders.Cardiovasc Toxicol. 2018 Dec;18(6):493-506. doi: 10.1007/s12012-018-9462-2. Cardiovasc Toxicol. 2018. PMID: 29968072 Review.
-
Efficacy and safety of poly (ADP-ribose) polymerase inhibitors therapy for BRCA-mutated breast cancer: A systematic review and meta-analysis.J Cancer Res Ther. 2021 Dec;17(7):1672-1678. doi: 10.4103/jcrt.jcrt_2085_21. J Cancer Res Ther. 2021. PMID: 35381738
Cited by
-
Nano-Zirconium Dioxide Catalyzed Multicomponent Synthesis of Bioactive Pyranopyrazoles That Target Cyclin Dependent Kinase 1 in Human Breast Cancer Cells.Biomedicines. 2023 Jan 10;11(1):172. doi: 10.3390/biomedicines11010172. Biomedicines. 2023. PMID: 36672680 Free PMC article.
-
Discovery of oxazine-linked pyrimidine as an inhibitor of breast cancer growth and metastasis by abrogating NF-κB activation.Front Oncol. 2024 Jul 29;14:1390992. doi: 10.3389/fonc.2024.1390992. eCollection 2024. Front Oncol. 2024. PMID: 39135991 Free PMC article.
-
Repurposing Study of 4-Acyl-1-phenylaminocarbonyl-2-substituted-piperazine Derivatives as Potential Anticancer Agents-In Vitro Evaluation against Breast Cancer Cells.Int J Mol Sci. 2023 Dec 1;24(23):17041. doi: 10.3390/ijms242317041. Int J Mol Sci. 2023. PMID: 38069364 Free PMC article.
-
Discovery of Pyrimidine- and Coumarin-Linked Hybrid Molecules as Inducers of JNK Phosphorylation through ROS Generation in Breast Cancer Cells.Molecules. 2023 Apr 13;28(8):3450. doi: 10.3390/molecules28083450. Molecules. 2023. PMID: 37110684 Free PMC article.
-
Development of Piperazine- and Oxazine-Linked Pyrimidines as p65 Subunit Binders of NF-κB in Human Breast Cancer Cells.Biomedicines. 2023 Oct 6;11(10):2716. doi: 10.3390/biomedicines11102716. Biomedicines. 2023. PMID: 37893090 Free PMC article.
References
-
- Oza A.M., Tinker A.V., Oaknin A., Shapira-Frommer R., McNeish I.A., Swisher E.M., Ray-Coquard I., Bell-McGuinn K., Coleman R.L., O’Malley D.M., et al. Antitumor activity and safety of the PARP inhibitor rucaparib in patients with high-grade ovarian carcinoma and a germline or somatic BRCA1 or BRCA2 mutation: Integrated analysis of data from study 10 and ARIEL2. Gynecol. Oncol. 2017;147:267–275. doi: 10.1016/j.ygyno.2017.08.022. - DOI - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

