Predicting clinical scores in Huntington's disease: a lightweight speech test
- PMID: 35567614
- PMCID: PMC9363375
- DOI: 10.1007/s00415-022-11148-1
Predicting clinical scores in Huntington's disease: a lightweight speech test
Abstract
Objectives: Using brief samples of speech recordings, we aimed at predicting, through machine learning, the clinical performance in Huntington's Disease (HD), an inherited Neurodegenerative disease (NDD).
Methods: We collected and analyzed 126 samples of audio recordings of both forward and backward counting from 103 Huntington's disease gene carriers [87 manifest and 16 premanifest; mean age 50.6 (SD 11.2), range (27-88) years] from three multicenter prospective studies in France and Belgium (MIG-HD (ClinicalTrials.gov NCT00190450); BIO-HD (ClinicalTrials.gov NCT00190450) and Repair-HD (ClinicalTrials.gov NCT00190450). We pre-registered all of our methods before running any analyses, in order to avoid inflated results. We automatically extracted 60 speech features from blindly annotated samples. We used machine learning models to combine multiple speech features in order to make predictions at individual levels of the clinical markers. We trained machine learning models on 86% of the samples, the remaining 14% constituted the independent test set. We combined speech features with demographics variables (age, sex, CAG repeats, and burden score) to predict cognitive, motor, and functional scores of the Unified Huntington's disease rating scale. We provided correlation between speech variables and striatal volumes.
Results: Speech features combined with demographics allowed the prediction of the individual cognitive, motor, and functional scores with a relative error from 12.7 to 20.0% which is better than predictions using demographics and genetic information. Both mean and standard deviation of pause durations during backward recitation and clinical scores correlated with striatal atrophy (Spearman 0.6 and 0.5-0.6, respectively).
Interpretation: Brief and examiner-free speech recording and analysis may become in the future an efficient method for remote evaluation of the individual condition in HD and likely in other NDD.
Keywords: Huntington’s disease; Machine learning; Speech.
© 2022. The Author(s).
Conflict of interest statement
Nothing to report.
Figures




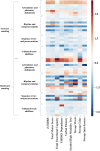
References
-
- (1996) Unified Huntington’s disease rating scale: reliability and consistency. Huntington study group. Mov Disord Off J Mov Disord. Soc 11(2): 136–142. 10.1002/mds.870110204. - PubMed
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials

