Statistic Copolymers Working as Growth Factor-Binding Mimics of Fibronectin
- PMID: 35570405
- PMCID: PMC9313494
- DOI: 10.1002/advs.202200775
Statistic Copolymers Working as Growth Factor-Binding Mimics of Fibronectin
Abstract
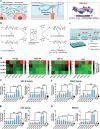
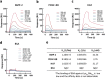
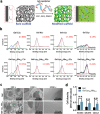
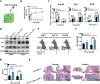
Growth factors (GFs) play important roles in biological system and are widely used in tissue regeneration. However, their application is greatly hindered by short in vivo lifetime of GFs. GFs are bound to fibronectin dynamically in the extracellular matrix, which inspired the authors to mimic the GF binding domain of fibronectin and design GF-binding amphiphilic copolymers bearing positive charges. The optimal amino acid polymer can bind to a variety of representative GFs, such as bone morphogenetic protein-2 (BMP-2) and TGF-β1 from the transforming growth factor-β superfamily, PDGF-AA and PDGF-BB from the platelet-derived growth factor family, FGF-10 and FGF-21 from the fibroblast growth factor family, epidermal growth factor from the EGF family and hepatocyte growth factor from the plasminogen-related growth factor family, with binding affinities up to the nanomolar level. 3D scaffolds immobilized with the optimal copolymer enable sustained release of loaded BMP-2 without burst release and significantly enhances the in vivo function of BMP-2 for bone formation. This strategy opens new avenues in designing GF-binding copolymers as synthetic mimics of fibronectin for diverse applications.
Keywords: fibronectin mimicking; growth factor binding; statistic copolymers.
© 2022 The Authors. Advanced Science published by Wiley-VCH GmbH.
Conflict of interest statement
R.L. and W.Z. are co‐inventors on a patent application covering reported materials and application to bind growth factors. All remaining authors declare no competing interests.
Figures

Similar articles
-
The 12th-14th type III repeats of fibronectin function as a highly promiscuous growth factor-binding domain.FASEB J. 2010 Dec;24(12):4711-21. doi: 10.1096/fj.09-151282. Epub 2010 Jul 29. FASEB J. 2010. PMID: 20671107
-
Human corneal fibroblast migration and extracellular matrix synthesis during stromal repair: Role played by platelet-derived growth factor-BB, basic fibroblast growth factor, and transforming growth factor-β1.J Tissue Eng Regen Med. 2018 Feb;12(2):e737-e746. doi: 10.1002/term.2360. Epub 2017 May 12. J Tissue Eng Regen Med. 2018. PMID: 27860426
-
Material-driven fibronectin assembly for high-efficiency presentation of growth factors.Sci Adv. 2016 Aug 26;2(8):e1600188. doi: 10.1126/sciadv.1600188. eCollection 2016 Aug. Sci Adv. 2016. PMID: 27574702 Free PMC article.
-
Preparation and Characterization of Human Adipose Tissue-Derived Extracellular Matrix, Growth Factors, and Stem Cells: A Concise Review.Tissue Eng Regen Med. 2019 Jul 5;16(4):385-393. doi: 10.1007/s13770-019-00199-7. eCollection 2019 Aug. Tissue Eng Regen Med. 2019. PMID: 31413942 Free PMC article. Review.
-
Growth factors in the treatment of Achilles tendon injury.Front Bioeng Biotechnol. 2023 Sep 14;11:1250533. doi: 10.3389/fbioe.2023.1250533. eCollection 2023. Front Bioeng Biotechnol. 2023. PMID: 37781529 Free PMC article. Review.
Cited by
-
Single-Center Trifunctional Organocatalyst Enables Fast and Controlled Polymerization on N-Carboxyanhydride.ACS Cent Sci. 2024 Oct 29;11(3):382-392. doi: 10.1021/acscentsci.4c01346. eCollection 2025 Mar 26. ACS Cent Sci. 2024. PMID: 40161954 Free PMC article.
-
Injectable composite microspheres/hydrogel membranes for Achilles tendon regeneration.Mater Today Bio. 2025 Jul 23;34:102129. doi: 10.1016/j.mtbio.2025.102129. eCollection 2025 Oct. Mater Today Bio. 2025. PMID: 40755897 Free PMC article.
-
An Off-the-Shelf Artificial Blood Clot Hydrogel Neutralizing Multiple Proinflammatory Mediators for Pro-Regenerative Periodontitis Treatment.Adv Sci (Weinh). 2025 Aug;12(31):e04106. doi: 10.1002/advs.202504106. Epub 2025 May 28. Adv Sci (Weinh). 2025. PMID: 40433947 Free PMC article.
-
Bioactive peptides and proteins for tissue repair: microenvironment modulation, rational delivery, and clinical potential.Mil Med Res. 2024 Dec 5;11(1):75. doi: 10.1186/s40779-024-00576-x. Mil Med Res. 2024. PMID: 39639374 Free PMC article. Review.
References
-
- Rapraeger A. C., Krufka A., Olwin B. B., Science 1991, 252, 1705. - PubMed
-
- a) Ehrbar M., Djonov V. G., Schnell C., Tschanz S. A., Martiny‐Baron G., Schenk U., Wood J., Burri P. H., Hubbell J. A., Zisch A. H., Circ. Res. 2004, 94, 1124; - PubMed
- b) Puccinelli T. J., Bertics P. J., Masters K. S., Acta Biomater. 2010, 6, 3415; - PMC - PubMed
- c) Hudalla G. A., Koepsel J. T., Murphy W. L., Adv. Mater. 2011, 23, 5415; - PMC - PubMed
- d) Chu H., Gao J., Chen C. W., Huard J., Wang Y., Proc. Natl. Acad. Sci. USA 2011, 108, 13444; - PMC - PubMed
- e) Samorezov J. E., Alsberg E., Adv. Drug Del. Rev. 2015, 84, 45; - PMC - PubMed
- f) Schwab E. H., Pohl T. L. M., Haraszti T., Schwaerzer G. K., Hiepen C., Spatz J. P., Knaus P., Cavalcanti‐Adam E. A., Nano Lett. 2015, 15, 1526; - PubMed
- g) Seo B.‐B., Koh J.‐T., Song S.‐C., Biomaterials 2017, 122, 91; - PubMed
- h) Tabisz B., Schmitz W., Schmitz M., Luehmann T., Heusler E., Rybak J.‐C., Meinel L., Fiebig J. E., Mueller T. D., Nickel J., Biomacromolecules 2017, 18, 695; - PubMed
- i) Andrew C. D., Pierluca P., Jessica N., Gráinne M. C., Daniel J. K., Biomaterials 2018, 162, 34. - PubMed
-
- Macri L., Silverstein D., Clark R. A. F., Adv. Drug Del. Rev. 2007, 59, 1366. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- 51621002/National Natural Science Foundation of China
- 31800801/National Natural Science Foundation of China
- 20XD1421400/Program of Shanghai Academic/Technology Research Leader
- 2021 Sci &/Shanghai Frontier Science Research Base of Optogenetic Techniques for Cell Metabolism (Shanghai Municipal Education Commission)
- Tech 03-28/Shanghai Frontier Science Research Base of Optogenetic Techniques for Cell Metabolism (Shanghai Municipal Education Commission)
LinkOut - more resources
Full Text Sources
