Structural anatomy of Protein Kinase C C1 domain interactions with diacylglycerol and other agonists
- PMID: 35577811
- PMCID: PMC9110374
- DOI: 10.1038/s41467-022-30389-2
Structural anatomy of Protein Kinase C C1 domain interactions with diacylglycerol and other agonists
Abstract
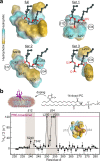
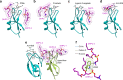
Diacylglycerol (DAG) is a versatile lipid whose 1,2-sn-stereoisomer serves both as second messenger in signal transduction pathways that control vital cellular processes, and as metabolic precursor for downstream signaling lipids such as phosphatidic acid. Effector proteins translocate to available DAG pools in the membranes by using conserved homology 1 (C1) domains as DAG-sensing modules. Yet, how C1 domains recognize and capture DAG in the complex environment of a biological membrane has remained unresolved for the 40 years since the discovery of Protein Kinase C (PKC) as the first member of the DAG effector cohort. Herein, we report the high-resolution crystal structures of a C1 domain (C1B from PKCδ) complexed to DAG and to each of four potent PKC agonists that produce different biological readouts and that command intense therapeutic interest. This structural information details the mechanisms of stereospecific recognition of DAG by the C1 domains, the functional properties of the lipid-binding site, and the identities of the key residues required for the recognition and capture of DAG and exogenous agonists. Moreover, the structures of the five C1 domain complexes provide the high-resolution guides for the design of agents that modulate the activities of DAG effector proteins.
© 2022. The Author(s).
Conflict of interest statement
S.S.K. and T.I.I. declare the existence of a potential financial interest due to a provisional patent application (Application No. 63/239,986) pending with the United States Patent and Trademark Office, with the specific aspect of this manuscript covered in the application being the method of crystallization of C1 domains complexed to ligands (T.I.I. and S.S.K. are listed as inventors and applicants, under obligation to assign to Texas A&M University). The remaining authors declare no competing interests.
Figures




Similar articles
-
Mechanism of diacylglycerol-induced membrane targeting and activation of protein kinase Cdelta.J Biol Chem. 2004 Jul 9;279(28):29501-12. doi: 10.1074/jbc.M403191200. Epub 2004 Apr 22. J Biol Chem. 2004. PMID: 15105418
-
Diacylglycerol-induced membrane targeting and activation of protein kinase Cepsilon: mechanistic differences between protein kinases Cdelta and Cepsilon.J Biol Chem. 2005 May 20;280(20):19784-93. doi: 10.1074/jbc.M411285200. Epub 2005 Mar 15. J Biol Chem. 2005. PMID: 15769752
-
C1 domains exposed: from diacylglycerol binding to protein-protein interactions.Biochim Biophys Acta. 2006 Aug;1761(8):827-37. doi: 10.1016/j.bbalip.2006.05.001. Epub 2006 May 13. Biochim Biophys Acta. 2006. PMID: 16861033 Review.
-
Interfacial partitioning of a loop hinge residue contributes to diacylglycerol affinity of conserved region 1 domains.J Biol Chem. 2014 Oct 3;289(40):27653-64. doi: 10.1074/jbc.M114.585570. Epub 2014 Aug 14. J Biol Chem. 2014. PMID: 25124034 Free PMC article.
-
Synthetic diacylglycerols (DAG) and DAG-lactones as activators of protein kinase C (PK-C).Acc Chem Res. 2003 Jun;36(6):434-43. doi: 10.1021/ar020124b. Acc Chem Res. 2003. PMID: 12809530 Review.
Cited by
-
An evolving view of complex II-noncanonical complexes, megacomplexes, respiration, signaling, and beyond.J Biol Chem. 2023 Jun;299(6):104761. doi: 10.1016/j.jbc.2023.104761. Epub 2023 Apr 27. J Biol Chem. 2023. PMID: 37119852 Free PMC article. Review.
-
Acute diacylglycerol production activates critical membrane-shaping proteins leading to mitochondrial tubulation and fission.Nat Commun. 2025 Mar 19;16(1):2685. doi: 10.1038/s41467-025-57439-9. Nat Commun. 2025. PMID: 40102394 Free PMC article.
-
Single-residue mutation in protein kinase C toggles between cancer and neurodegeneration.Biochem J. 2023 Aug 30;480(16):1299-1316. doi: 10.1042/BCJ20220397. Biochem J. 2023. PMID: 37551632 Free PMC article.
-
Ultrasmall chemogenetic tags with group-transfer ligands.bioRxiv [Preprint]. 2025 May 10:2025.05.10.653252. doi: 10.1101/2025.05.10.653252. bioRxiv. 2025. PMID: 40654995 Free PMC article. Preprint.
-
Crosstalk between diacylglycerol kinase and protein kinase A in the regulation of airway smooth muscle cell proliferation.Respir Res. 2023 Jun 10;24(1):155. doi: 10.1186/s12931-023-02465-8. Respir Res. 2023. PMID: 37301818 Free PMC article.
References
-
- Rozengurt E. Regulation and Function of Protein Kinase D Signaling. In: Protein Kinase C in Cancer Signaling and Therapy (ed Kazanietz MG). 117–154. (Humana Press, 2010).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

