Lineage-specific insertions in T-box riboswitches modulate antibiotic binding and action
- PMID: 35580054
- PMCID: PMC9177973
- DOI: 10.1093/nar/gkac359
Lineage-specific insertions in T-box riboswitches modulate antibiotic binding and action
Abstract
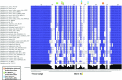
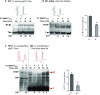
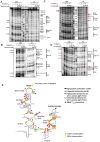
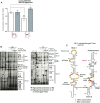
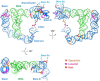
T-box riboswitches (T-boxes) are essential RNA regulatory elements with a remarkable structural diversity, especially among bacterial pathogens. In staphylococci, all glyS T-boxes synchronize glycine supply during synthesis of nascent polypeptides and cell wall formation and are characterized by a conserved and unique insertion in their antiterminator/terminator domain, termed stem Sa. Interestingly, in Staphylococcus aureus the stem Sa can accommodate binding of specific antibiotics, which in turn induce robust and diverse effects on T-box-mediated transcription. In the present study, domain swap mutagenesis and probing analysis were performed to decipher the role of stem Sa. Deletion of stem Sa significantly reduces both the S. aureus glyS T-box-mediated transcription readthrough levels and the ability to discriminate among tRNAGly isoacceptors, both in vitro and in vivo. Moreover, the deletion inverted the previously reported stimulatory effects of specific antibiotics. Interestingly, stem Sa insertion in the terminator/antiterminator domain of Geobacillus kaustophilus glyS T-box, which lacks this domain, resulted in elevated transcription in the presence of tigecycline and facilitated discrimination among proteinogenic and nonproteinogenic tRNAGly isoacceptors. Overall, stem Sa represents a lineage-specific structural feature required for efficient staphylococcal glyS T-box-mediated transcription and it could serve as a species-selective druggable target through its ability to modulate antibiotic binding.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures


Similar articles
-
A glyS T-box riboswitch with species-specific structural features responding to both proteinogenic and nonproteinogenic tRNAGly isoacceptors.RNA. 2015 Oct;21(10):1790-806. doi: 10.1261/rna.052712.115. Epub 2015 Aug 14. RNA. 2015. PMID: 26276802 Free PMC article.
-
Direct modulation of T-box riboswitch-controlled transcription by protein synthesis inhibitors.Nucleic Acids Res. 2017 Sep 29;45(17):10242-10258. doi: 10.1093/nar/gkx663. Nucleic Acids Res. 2017. PMID: 28973457 Free PMC article.
-
An evolving tale of two interacting RNAs-themes and variations of the T-box riboswitch mechanism.IUBMB Life. 2019 Aug;71(8):1167-1180. doi: 10.1002/iub.2098. Epub 2019 Jun 17. IUBMB Life. 2019. PMID: 31206978 Free PMC article. Review.
-
Methionine biosynthesis in Staphylococcus aureus is tightly controlled by a hierarchical network involving an initiator tRNA-specific T-box riboswitch.PLoS Pathog. 2013 Sep;9(9):e1003606. doi: 10.1371/journal.ppat.1003606. Epub 2013 Sep 12. PLoS Pathog. 2013. PMID: 24068926 Free PMC article.
-
Riboswitches: discovery of drugs that target bacterial gene-regulatory RNAs.Acc Chem Res. 2011 Dec 20;44(12):1329-38. doi: 10.1021/ar200039b. Epub 2011 May 26. Acc Chem Res. 2011. PMID: 21615107 Free PMC article. Review.
Cited by
-
Observation of SAM-VI Riboswitch Dynamics Using Single-Molecule FRET.Biomolecules. 2025 Jun 9;15(6):841. doi: 10.3390/biom15060841. Biomolecules. 2025. PMID: 40563481 Free PMC article.
-
Structural idiosyncrasies of glycyl T-box riboswitches among pathogenic bacteria.RNA. 2024 Sep 16;30(10):1328-1344. doi: 10.1261/rna.080071.124. RNA. 2024. PMID: 38981655
-
Coupling tRNAGly gene redundancy with staphylococcal cell wall integrity, antibiotic susceptibility, and virulence potential.Nucleic Acids Res. 2025 Jul 8;53(13):gkaf599. doi: 10.1093/nar/gkaf599. Nucleic Acids Res. 2025. PMID: 40653316 Free PMC article.
-
A Riboswitch-Driven Era of New Antibacterials.Antibiotics (Basel). 2022 Sep 13;11(9):1243. doi: 10.3390/antibiotics11091243. Antibiotics (Basel). 2022. PMID: 36140022 Free PMC article. Review.
-
Observation of structural switch in nascent SAM-VI riboswitch during transcription at single-nucleotide and single-molecule resolution.Nat Commun. 2023 Apr 22;14(1):2320. doi: 10.1038/s41467-023-38042-2. Nat Commun. 2023. PMID: 37087479 Free PMC article.
References
-
- Grundy F.J., Henkin T.M.. tRNA as a positive regulator of transcription antitermination in B. subtilis. Cell. 1993; 74:475–482. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials

