In Vitro Evolution of Listeria monocytogenes Reveals Selective Pressure for Loss of SigB and AgrA Function at Different Incubation Temperatures
- PMID: 35583325
- PMCID: PMC9195950
- DOI: 10.1128/aem.00330-22
In Vitro Evolution of Listeria monocytogenes Reveals Selective Pressure for Loss of SigB and AgrA Function at Different Incubation Temperatures
Abstract
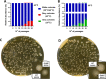
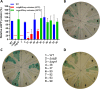
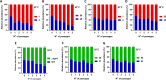
The alternative sigma factor B (σB) contributes to the stress tolerance of the foodborne pathogen Listeria monocytogenes by upregulating the general stress response. We previously showed that σB loss-of-function mutations arise frequently in strains of L. monocytogenes and suggested that mild stresses might favor the selection of such mutations. In this study, we performed in vitro evolution experiments (IVEE) where L. monocytogenes was allowed to evolve over 30 days at elevated (42°C) or lower (30°C) incubation temperatures. Isolates purified throughout the IVEE revealed the emergence of sigB operon mutations at 42°C. However, at 30°C, independent alleles in the agr locus arose, resulting in the inactivation of Agr quorum sensing. Colonies of both sigB mutants and agr mutants exhibited a greyer coloration on 7-days-old agar plates than those of the parental strain. Scanning electron microscopy revealed a more complex colony architecture in the wild type than in the mutant strains. sigB mutant strains outcompeted the parental strain at 42°C but not at 30°C, while agr mutant strains showed a small increase in competitive fitness at 30°C. Analysis of 40,080 L. monocytogenes publicly available genome sequences revealed a high occurrence rate of premature stop codons in both the sigB and agrCA loci. An analysis of a local L. monocytogenes strain collection revealed 5 out of 168 strains carrying agrCA alleles. Our results suggest that the loss of σB or Agr confer an increased competitive fitness in some specific conditions and this likely contributes to the emergence of these alleles in strains of L. monocytogenes. IMPORTANCE To withstand environmental aggressions, L. monocytogenes upregulates a large regulon through the action of the alternative sigma factor B (σB). However, σB becomes detrimental for L. monocytogenes growth under mild stresses, which confer a competitive advantage to σB loss-of-function alleles. Temperatures of 42°C, a mild stress, are often employed in mutagenesis protocols of L. monocytogenes and promote the emergence of σB loss-of-function alleles in the sigB operon. In contrast, lower temperatures of 30°C promote the emergence of Agr loss-of-function alleles, a cell-cell communication mechanism in L. monocytogenes. Our findings demonstrate that loss-of-function alleles emerge spontaneously in laboratory-grown strains. These alleles rise in the population as a consequence of the trade-off between growth and survival imposed by the activation of σB in L. monocytogenes. Additionally, our results demonstrate the importance of identifying unwanted hitchhiker mutations in newly constructed mutant strains.
Keywords: Listeria monocytogenes; agr system; bacterial competition; general stress response; in vitro evolution; quorum sensing; σB.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




References
-
- World Health Organization. 2018. Listeriosis. World Health Organization, Geneva, Switzerland. https://www.who.int/news-room/fact-sheets/detail/listeriosis. Accessed 2 June 2021.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

