Zn-regulated GTPase metalloprotein activator 1 modulates vertebrate zinc homeostasis
- PMID: 35584702
- PMCID: PMC9189065
- DOI: 10.1016/j.cell.2022.04.011
Zn-regulated GTPase metalloprotein activator 1 modulates vertebrate zinc homeostasis
Abstract
Zinc (Zn) is an essential micronutrient and cofactor for up to 10% of proteins in living organisms. During Zn limitation, specialized enzymes called metallochaperones are predicted to allocate Zn to specific metalloproteins. This function has been putatively assigned to G3E GTPase COG0523 proteins, yet no Zn metallochaperone has been experimentally identified in any organism. Here, we functionally characterize a family of COG0523 proteins that is conserved across vertebrates. We identify Zn metalloprotease methionine aminopeptidase 1 (METAP1) as a COG0523 client, leading to the redesignation of this group of COG0523 proteins as the Zn-regulated GTPase metalloprotein activator (ZNG1) family. Using biochemical, structural, genetic, and pharmacological approaches across evolutionarily divergent models, including zebrafish and mice, we demonstrate a critical role for ZNG1 proteins in regulating cellular Zn homeostasis. Collectively, these data reveal the existence of a family of Zn metallochaperones and assign ZNG1 an important role for intracellular Zn trafficking.
Keywords: CBWD; COG0523; GTPase; METAP1; ZNG1; metallochaperone; metalloprotein; zf-C6H2; zinc; zinc finger.
Copyright © 2022 Elsevier Inc. All rights reserved.
Conflict of interest statement
Declaration of interests The authors declare no competing interests.
Figures






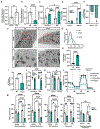
Comment in
-
A zinc chaperone mediates the flow of an inorganic commodity to an important cellular client.Cell. 2022 Jun 9;185(12):2013-2015. doi: 10.1016/j.cell.2022.05.012. Cell. 2022. PMID: 35688131
-
Escort proteins for cellular zinc ions.Nature. 2022 Aug;608(7921):38-39. doi: 10.1038/d41586-022-01988-2. Nature. 2022. PMID: 35918525 No abstract available.
References
-
- Addlagatta A, Hu X, Liu JO, and Matthews BW (2005). Structural basis for the functional differences between type I and type II human methionine aminopeptidases. Biochemistry 44, 14741–14749. - PubMed
-
- Andreini C, Banci L, Bertini I, and Rosato A (2006). Counting the zinc-proteins encoded in the human genome. J Proteome Res 5, 196–201. - PubMed
-
- Arfin SM, and Bradshaw RA (1988). Cotranslational processing and protein turnover in eukaryotic cells. Biochemistry 27, 7979–7984. - PubMed
-
- Ba LA, Doering M, Burkholz T, and Jacob C (2009). Metal trafficking: from maintaining the metal homeostasis to future drug design. Metallomics 1, 292–311. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
- P30 DK058404/DK/NIDDK NIH HHS/United States
- R01 AI145992/AI/NIAID NIH HHS/United States
- F32 GM142246/GM/NIGMS NIH HHS/United States
- T32 GM109825/GM/NIGMS NIH HHS/United States
- T32 DK101003/DK/NIDDK NIH HHS/United States
- P30 EY008126/EY/NEI NIH HHS/United States
- R01 AI138581/AI/NIAID NIH HHS/United States
- F32 HL144081/HL/NHLBI NIH HHS/United States
- T32 GM131994/GM/NIGMS NIH HHS/United States
- U24 DK059637/DK/NIDDK NIH HHS/United States
- T32 ES007028/ES/NIEHS NIH HHS/United States
- R01 AI101171/AI/NIAID NIH HHS/United States
- F32 AI157215/AI/NIAID NIH HHS/United States
- P30 DK020593/DK/NIDDK NIH HHS/United States
- R01 AI150701/AI/NIAID NIH HHS/United States
- P30 CA068485/CA/NCI NIH HHS/United States
- R35 GM118157/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

