Prognosis prediction of stage IV colorectal cancer patients by mRNA transcriptional profile
- PMID: 35587572
- PMCID: PMC9761091
- DOI: 10.1002/cam4.4824
Prognosis prediction of stage IV colorectal cancer patients by mRNA transcriptional profile
Abstract
Background: Stage IV colorectal cancer patients with liver metastasis represent a special group of CRC patients with poor prognosis. The prognostic factors have not been investigated for stage IV CRC patients undergoing primary cancer resection but not candidates for metastasis resection.
Methods: Ninety-nine stage IV CRC patients who underwent primary cancer resection without metastasis resection were retrospectively recruited. Both whole-exome sequencing (WES) and RNA-seq were performed with frozen primary cancer tissues, using para-cancerous normal tissues as the control. Valid data were obtained from 78 patients for WES and 84 patients for RNA-seq. Univariate, multivariate Cox analyses were performed and Nomogram model was established to predict patient prognosis.
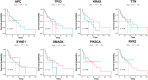
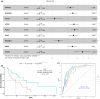
Results: The correlation between patient prognosis and clinicopathological factors, mutational status, or mRNA level changes was examined. Univariate (p = 0.0007) and subsequent multivariate analyses on clinicopathological factors showed that location (left or right) was the only independent risk factor for patient prognosis (HR = 3.63; 95% CI: 1.56-8.40, p = 0.003), while T, N, M staging, gender, race, location (rectum or colon), and pathological types were not stratifying factors. The mutational status of APC, TP53, KRAS, TTN, SYNE1, SMAD4, PIK3CA, RYR2, and BRAF did not show significant stratification in patient prognosis. RNA-seq showed that genes related to membrane function, ion channels, transporters, or receptors were among those with significant mRNA level alterations. Univariate analysis identified 97 genes with significantly altered mRNA levels, while NEUROD1, FGF18, SFTA2, PLAC1, SAA2, DSCAML1, and OTOP3 were significant in multivariate analysis. A risk model was established to stratify the prognosis of stage IV CRC patients. A Nomogram model was established with these genes to predict individual patient prognosis.
Conclusions: A panel of eight genes with significant mRNA level alterations was capable of predicting the prognosis and risk of the specific patient group. Future prospective study is needed to validate the model.
Keywords: RNA-seq; WES; colorectal cancer; expression; mutation; prognosis; transcription.
© 2022 The Authors. Cancer Medicine published by John Wiley & Sons Ltd.
Conflict of interest statement
All authors claim no conflict of interest.
Figures





References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

