Disruption of Toxoplasma gondii-Induced Host Cell DNA Replication Is Dependent on Contact Inhibition and Host Cell Type
- PMID: 35587658
- PMCID: PMC9241542
- DOI: 10.1128/msphere.00160-22
Disruption of Toxoplasma gondii-Induced Host Cell DNA Replication Is Dependent on Contact Inhibition and Host Cell Type
Abstract
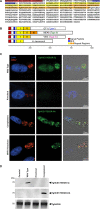
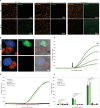
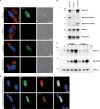
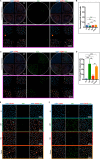
The protozoan Toxoplasma gondii is a highly successful obligate intracellular parasite that, upon invasion of its host cell, releases an array of host-modulating protein effectors to counter host defenses and further its own replication and dissemination. Early studies investigating the impact of T. gondii infection on host cell function revealed that this parasite can force normally quiescent cells to activate their cell cycle program. Prior reports by two independent groups identified the dense granule protein effector HCE1/TEEGR as being solely responsible for driving host cell transcriptional changes through its direct interaction with the cyclin E regulatory complex DP1 and associated transcription factors. Our group independently identified HCE1/TEEGR through the presence of distinct repeated regions found in a number of host nuclear targeted parasite effectors and verified its central role in initiating host cell cycle changes. Additionally, we report here the time-resolved kinetics of host cell cycle transition in response to HCE1/TEEGR, using the fluorescence ubiquitination cell cycle indicator reporter line (FUCCI), and reveal the existence of a block in S-phase progression and host DNA synthesis in several cell lines commonly used in the study of T. gondii. Importantly, we have observed that this S-phase block is not due to additional dense granule effectors but rather is dependent on the host cell line background and contact inhibition status of the host monolayer in vitro. This work highlights intriguing differences in the host response to reprogramming by the parasite and raises interesting questions regarding how parasite effectors differentially manipulate the host cell depending on the in vitro or in vivo context. IMPORTANCE Toxoplasma gondii chronically infects approximately one-third of the global population and can produce severe pathology in immunologically immature or compromised individuals. During infection, this parasite releases numerous host-targeted effector proteins that can dramatically alter the expression of a variety of host genes. A better understanding of parasite effectors and their host targets has the potential to not only provide ways to control infection but also inform us about our own basic biology. One host pathway that has been known to be altered by T. gondii infection is the cell cycle, and prior reports have identified a parasite effector, known as HCE1/TEEGR, as being responsible. In this report, we further our understanding of the kinetics of cell cycle transition induced by this effector and show that the capacity of HCE1/TEEGR to induce host cell DNA synthesis is dependent on both the cell type and the status of contact inhibition.
Keywords: FUCCI; HCE1; HCE1/TEEGR; S-phase; TEEGR; Toxoplasma gondii; cell cycle; cyclin E; host-parasite interaction; parasite effectors.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


References
-
- Dubey JP, Beattie CP. 1988. Toxoplasmosis of animals and man. CRC Press, Boca Raton, FL.
-
- Dubey JP. 2007. The history and life cycle of Toxoplasma gondii, p 1–17. In Weiss LM, Kim K (ed), Toxoplasma gondii, the model Apicomplexan: perspectives and methods. Elsevier, New York, NY.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
