Evolution and functional diversification of R2R3-MYB transcription factors in plants
- PMID: 35591925
- PMCID: PMC9113232
- DOI: 10.1093/hr/uhac058
Evolution and functional diversification of R2R3-MYB transcription factors in plants
Abstract
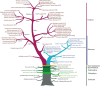
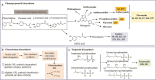
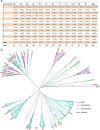
R2R3-MYB genes (R2R3-MYBs) form one of the largest transcription factor gene families in the plant kingdom, with substantial structural and functional diversity. However, the evolutionary processes leading to this amazing functional diversity have not yet been clearly established. Recently developed genomic and classical molecular technologies have provided detailed insights into the evolutionary relationships and functions of plant R2R3-MYBs. Here, we review recent genome-level and functional analyses of plant R2R3-MYBs, with an emphasis on their evolution and functional diversification. In land plants, this gene family underwent a large expansion by whole genome duplications and small-scale duplications. Along with this population explosion, a series of functionally conserved or lineage-specific subfamilies/groups arose with roles in three major plant-specific biological processes: development and cell differentiation, specialized metabolism, and biotic and abiotic stresses. The rapid expansion and functional diversification of plant R2R3-MYBs are highly consistent with the increasing complexity of angiosperms. In particular, recently derived R2R3-MYBs with three highly homologous intron patterns (a, b, and c) are disproportionately related to specialized metabolism and have become the predominant subfamilies in land plant genomes. The evolution of plant R2R3-MYBs is an active area of research, and further studies are expected to improve our understanding of the evolution and functional diversification of this gene family.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nanjing Agricultural University.
Figures

Similar articles
-
In silico analysis of R2R3-MYB transcription factors in the basal eudicot model, Aquilegia coerulea.3 Biotech. 2024 Nov;14(11):284. doi: 10.1007/s13205-024-04119-y. Epub 2024 Oct 29. 3 Biotech. 2024. PMID: 39479299 Free PMC article.
-
Insights into the Diversification and Evolution of R2R3-MYB Transcription Factors in Plants.Plant Physiol. 2020 Jun;183(2):637-655. doi: 10.1104/pp.19.01082. Epub 2020 Apr 14. Plant Physiol. 2020. PMID: 32291329 Free PMC article.
-
Diversification of R2R3-MYB Transcription Factors in the Tomato Family Solanaceae.J Mol Evol. 2016 Aug;83(1-2):26-37. doi: 10.1007/s00239-016-9750-z. Epub 2016 Jun 30. J Mol Evol. 2016. PMID: 27364496
-
Conservation and diversification of flavonoid metabolism in the plant kingdom.Curr Opin Plant Biol. 2020 Jun;55:100-108. doi: 10.1016/j.pbi.2020.04.004. Epub 2020 May 15. Curr Opin Plant Biol. 2020. PMID: 32422532 Review.
-
Roles of R2R3-MYB transcription factors in transcriptional regulation of anthocyanin biosynthesis in horticultural plants.Plant Mol Biol. 2018 Sep;98(1-2):1-18. doi: 10.1007/s11103-018-0771-4. Epub 2018 Aug 30. Plant Mol Biol. 2018. PMID: 30167900 Review.
Cited by
-
OsMYB58 Negatively Regulates Plant Growth and Development by Regulating Phosphate Homeostasis.Int J Mol Sci. 2024 Feb 12;25(4):2209. doi: 10.3390/ijms25042209. Int J Mol Sci. 2024. PMID: 38396886 Free PMC article.
-
The genomic and bulked segregant analysis of Curcuma alismatifolia revealed its diverse bract pigmentation.aBIOTECH. 2022 Oct 6;3(3):178-196. doi: 10.1007/s42994-022-00081-6. eCollection 2022 Sep. aBIOTECH. 2022. PMID: 36304840 Free PMC article.
-
The characteristic analysis of TaTDF1 reveals its function related to male sterility in wheat (Triticum aestivum L.).BMC Plant Biol. 2024 Aug 5;24(1):746. doi: 10.1186/s12870-024-05456-z. BMC Plant Biol. 2024. PMID: 39098914 Free PMC article.
-
The Transcription Factor MYB37 Positively Regulates Photosynthetic Inhibition and Oxidative Damage in Arabidopsis Leaves Under Salt Stress.Front Plant Sci. 2022 Jul 12;13:943153. doi: 10.3389/fpls.2022.943153. eCollection 2022. Front Plant Sci. 2022. PMID: 35903240 Free PMC article.
-
Genome-Wide Identification and Analysis of R2R3-MYB Genes Response to Saline-Alkali Stress in Quinoa.Int J Mol Sci. 2023 May 23;24(11):9132. doi: 10.3390/ijms24119132. Int J Mol Sci. 2023. PMID: 37298082 Free PMC article.
References
-
- Klempnauer KH, Gonda TJ, Michael Bishop J. Nucleotide sequence of the retroviral leukemia gene v-MYB and its cellular progenitor c-MYB: the architecture of a transduced oncogene. Cell. 1982;31:453–63. - PubMed
-
- Lipsick JS. One billion years of Myb. Oncogene. 1996;13:223–35. - PubMed
-
- Rosinski JA, Atchley WR. Molecular evolution of the MYB family of transcription factors: evidence for polyphyletic origin. J Mol Evol. 1998;46:74–83. - PubMed
-
- Kranz H, Scholz K, Weisshaar B. C-MYB oncogene-like genes encoding three MYB repeats occur in all major plant lineages. Plant J. 2000;21:231–5. - PubMed
-
- Dubos C, Stracke R, Grotewold Eet al. . MYB transcription factors in Arabidopsis. Trends Plant Sci. 2010;15:573–81. - PubMed
LinkOut - more resources
Full Text Sources

