Alternative end-joining in BCR gene rearrangements and translocations
- PMID: 35593472
- PMCID: PMC9828324
- DOI: 10.3724/abbs.2022051
Alternative end-joining in BCR gene rearrangements and translocations
Abstract
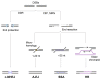
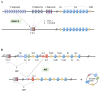
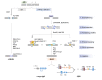
Programmed DNA double-strand breaks (DSBs) occur during antigen receptor gene recombination, namely V(D)J recombination in developing B lymphocytes and class switch recombination (CSR) in mature B cells. Repair of these DSBs by classical end-joining (c-NHEJ) enables the generation of diverse BCR repertoires for efficient humoral immunity. Deletion of or mutation in c-NHEJ genes in mice and humans confer various degrees of primary immune deficiency and predisposition to lymphoid malignancies that often harbor oncogenic chromosomal translocations. In the absence of c-NHEJ, alternative end-joining (A-EJ) catalyzes robust CSR and to a much lesser extent, V(D)J recombination, but the mechanisms of A-EJ are only poorly defined. In this review, we introduce recent advances in the understanding of A-EJ in the context of V(D)J recombination and CSR with emphases on DSB end processing, DNA polymerases and ligases, and discuss the implications of A-EJ to lymphoid development and chromosomal translocations.
Keywords: DSB end resection; V(D)J recombination; alternative end-joining; chromosomal translocation; classical nonhomologous end-joining; endonuclease.
Conflict of interest statement
The authors declare that they have no conflict of interest.
Figures
References
-
- Symington LS. Mechanism and regulation of DNA end resection in eukaryotes. Crit Rev Biochem Mol Biol. . 2016;51:195–212. doi: 10.3109/10409238.2016.1172552. - DOI - PMC - PubMed
-
- Chang HHY, Pannunzio NR, Adachi N, Lieber MR. Non-homologous DNA end joining and alternative pathways to double-strand break repair. Nat Rev Mol Cell Biol. . 2017;18:495–506. doi: 10.1038/nrm.2017.48. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

