Identification, Characterization and Function of Orphan Genes Among the Current Cucurbitaceae Genomes
- PMID: 35599909
- PMCID: PMC9114813
- DOI: 10.3389/fpls.2022.872137
Identification, Characterization and Function of Orphan Genes Among the Current Cucurbitaceae Genomes
Abstract
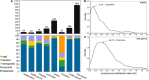
Orphan genes (OGs) that are missing identifiable homologs in other lineages may potentially make contributions to a variety of biological functions. The Cucurbitaceae family consists of a wide range of fruit crops of worldwide or local economic significance. To date, very few functional mechanisms of OGs in Cucurbitaceae are known. In this study, we systematically identified the OGs of eight Cucurbitaceae species using a comparative genomics approach. The content of OGs varied widely among the eight Cucurbitaceae species, ranging from 1.63% in chayote to 16.55% in wax gourd. Genetic structure analysis showed that OGs have significantly shorter protein lengths and fewer exons in Cucurbitaceae. The subcellular localizations of OGs were basically the same, with only subtle differences. Except for aggregation in some chromosomal regions, the distribution density of OGs was higher near the telomeres and relatively evenly distributed on the chromosomes. Gene expression analysis revealed that OGs had less abundantly and highly tissue-specific expression. Interestingly, the largest proportion of these OGs was significantly more tissue-specific expressed in the flower than in other tissues, and more detectable expression was found in the male flower. Functional prediction of OGs showed that (1) 18 OGs associated with male sterility in watermelon; (2) 182 OGs associated with flower development in cucumber; (3) 51 OGs associated with environmental adaptation in watermelon; (4) 520 OGs may help with the large fruit size in wax gourd. Our results provide the molecular basis and research direction for some important mechanisms in Cucurbitaceae species and domesticated crops.
Keywords: Cucurbitaceae; environmental adaptation; male sterility; orphan genes; transcriptome.
Copyright © 2022 Ma, Lai, Ding, Zhang, Chang, Li, Zhao and Zhong.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






Similar articles
-
The Lost and Found: Unraveling the Functions of Orphan Genes.J Dev Biol. 2023 Jun 13;11(2):27. doi: 10.3390/jdb11020027. J Dev Biol. 2023. PMID: 37367481 Free PMC article. Review.
-
Genome-Wide Identification, Characterization, and Expression Analysis of Orphan Genes Within Coriander.Plants (Basel). 2025 Mar 3;14(5):778. doi: 10.3390/plants14050778. Plants (Basel). 2025. PMID: 40094770 Free PMC article.
-
Genome-Wide Identification of YABBY Gene Family in Cucurbitaceae and Expression Analysis in Cucumber (Cucumis sativus L.).Genes (Basel). 2022 Mar 7;13(3):467. doi: 10.3390/genes13030467. Genes (Basel). 2022. PMID: 35328021 Free PMC article.
-
Orphan genes are involved in drought adaptations and ecoclimatic-oriented selections in domesticated cowpea.J Exp Bot. 2019 Jun 28;70(12):3101-3110. doi: 10.1093/jxb/erz145. J Exp Bot. 2019. PMID: 30949664
-
Genetic architecture of fruit size and shape variation in cucurbits: a comparative perspective.Theor Appl Genet. 2020 Jan;133(1):1-21. doi: 10.1007/s00122-019-03481-3. Epub 2019 Nov 25. Theor Appl Genet. 2020. PMID: 31768603 Review.
Cited by
-
Research Advances and Prospects of Orphan Genes in Plants.Front Plant Sci. 2022 Jul 8;13:947129. doi: 10.3389/fpls.2022.947129. eCollection 2022. Front Plant Sci. 2022. PMID: 35874010 Free PMC article. Review.
-
Genome-wide identification and expression analysis of orphan genes in twelve Musa (sub)species.3 Biotech. 2025 Feb;15(2):41. doi: 10.1007/s13205-025-04213-9. Epub 2025 Jan 14. 3 Biotech. 2025. PMID: 39822754
-
Orphan gene BR2 positively regulates bolting resistance through the vernalization pathway in Chinese cabbage.Hortic Res. 2024 Jul 30;11(10):uhae216. doi: 10.1093/hr/uhae216. eCollection 2024 Oct. Hortic Res. 2024. PMID: 39398948 Free PMC article.
-
The Lost and Found: Unraveling the Functions of Orphan Genes.J Dev Biol. 2023 Jun 13;11(2):27. doi: 10.3390/jdb11020027. J Dev Biol. 2023. PMID: 37367481 Free PMC article. Review.
-
Application of machine learning and genomics for orphan crop improvement.Nat Commun. 2025 Jan 24;16(1):982. doi: 10.1038/s41467-025-56330-x. Nat Commun. 2025. PMID: 39856113 Free PMC article. Review.
References
-
- Clément C., Audran J. C. (1995). Anther wall layers control pollen sugar nutrition in Lilium. Protoplasma 187 172–181. 10.1007/BF01280246 - DOI
LinkOut - more resources
Full Text Sources

