Interlaboratory Studies Using the NISTmAb to Advance Biopharmaceutical Structural Analytics
- PMID: 35601836
- PMCID: PMC9117750
- DOI: 10.3389/fmolb.2022.876780
Interlaboratory Studies Using the NISTmAb to Advance Biopharmaceutical Structural Analytics
Abstract
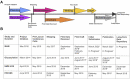
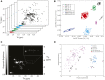
Biopharmaceuticals such as monoclonal antibodies are required to be rigorously characterized using a wide range of analytical methods. Various material properties must be characterized and well controlled to assure that clinically relevant features and critical quality attributes are maintained. A thorough understanding of analytical method performance metrics, particularly emerging methods designed to address measurement gaps, is required to assure methods are appropriate for their intended use in assuring drug safety, stability, and functional activity. To this end, a series of interlaboratory studies have been conducted using NISTmAb, a biopharmaceutical-representative and publicly available monoclonal antibody test material, to report on state-of-the-art method performance, harmonize best practices, and inform on potential gaps in the analytical measurement infrastructure. Reported here is a summary of the study designs, results, and future perspectives revealed from these interlaboratory studies which focused on primary structure, post-translational modifications, and higher order structure measurements currently employed during biopharmaceutical development.
Keywords: biopharmaceutical; interlaboratory study; monoclonal antibody; nistmab; therapeutic protein.
Copyright © 2022 Yandrofski, Mouchahoir, De Leoz, Duewer, Hudgens, Anderson, Arbogast, Delaglio, Brinson, Marino, Phinney, Tarlov and Schiel.
Conflict of interest statement
Author MLdL was employed by company Agilent Technologies. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



References
-
- Anderson K. W., Bergonzo C., Scott K., Karageorgos I. L., Gallagher E. S., Tayi V. S., et al. (2022). HDX-MS and MD Simulations Provide Evidence for Stabilization of the IgG1-FcγRIa (CD64a) Immune Complex through Intermolecular Glycoprotein Bonds. J. Mol. Biol. 434 (2), 167391. 10.1016/j.jmb.2021.167391 - DOI - PubMed
Publication types
LinkOut - more resources
Full Text Sources

