Beneficial commensal bacteria promote Drosophila growth by downregulating the expression of peptidoglycan recognition proteins
- PMID: 35601912
- PMCID: PMC9121327
- DOI: 10.1016/j.isci.2022.104357
Beneficial commensal bacteria promote Drosophila growth by downregulating the expression of peptidoglycan recognition proteins
Abstract
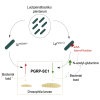
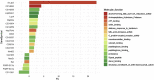
Commensal bacteria are known to promote host growth. Such effect partly relies on the capacity of microbes to regulate the host's transcriptional response. However, these evidences mainly come from comparing the transcriptional response caused by commensal bacteria with that of axenic animals, making it difficult to identify the animal genes that are specifically regulated by beneficial microbes. Here, we employ Drosophila melanogaster associated with Lactiplantibacillus plantarum to understand the host genetic pathways regulated by beneficial bacteria and leading to improved host growth. We show that microbial benefit to the host relies on the downregulation of peptidoglycan-recognition proteins. Specifically, we report that bacterial proliferation triggers the lower expression of PGRP-SC1 in larval midgut, which ultimately leads to improved host growth and development. Our study helps elucidate the mechanisms underlying the beneficial effect exerted by commensal bacteria, defining the role of immune effectors in the relationship between Drosophila and its gut microbes.
Keywords: immunology; microbiology; microbiome.
© 2022 The Author(s).
Conflict of interest statement
The authors declared no competing interests.
Figures



References
-
- Araújo A.R., Reis M., Rocha H., Aguiar B., Morales-Hojas R., Macedo-Ribeiro S., Fonseca N.A., Reboiro-Jato D., Reboiro-Jato M., Fdez-Riverola F., et al. The Drosophila melanogaster methuselah gene: a novel gene with ancient functions. PLoS One. 2013;8:1–12. doi: 10.1371/journal.pone.0063747. - DOI - PMC - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

