Litter Mixing Alters Microbial Decomposer Community to Accelerate Tomato Root Litter Decomposition
- PMID: 35604181
- PMCID: PMC9241821
- DOI: 10.1128/spectrum.00186-22
Litter Mixing Alters Microbial Decomposer Community to Accelerate Tomato Root Litter Decomposition
Abstract
Mixing plant litters of multiple species can alter litter decomposition, a key driver of carbon and nutrient cycling in terrestrial ecosystems. Changes in microbial decomposer communities is proposed as one of the mechanisms explaining this litter-mixture effect, but the underlying mechanism is unclear. In a microcosm litterbag experiment, we found that, at the early stage of decomposition, litter mixing promoted tomato root litter decomposition, thus generating a synergistic nonadditive litter-mixture effect. The transplanting decomposer community experiment showed that changes in microbial decomposer communities contributed to the nonadditive litter-mixture effect on tomato root litter decomposition. Moreover, litter mixing altered the abundance and diversity of bacterial and fungal communities on tomato root litter. Litter mixing also stimulated several putative keystone operational taxonomic units (OTUs) in the microbial correlation network, such as Fusarium sp. fOTU761 and Microbacterium sp. bOTU6632. Then, we isolated and cultured representative isolates of these two taxa, named Fusarium sp. F13 and Microbacterium sp. B26. Subsequent in vitro tests found that F13, but not B26, had strong decomposing ability; moreover, these two isolates developed synergistic interaction, thus promoted litter decomposition in coculture. Addition of F13 or B26 both promoted the decomposing activity of the resident decomposer community on tomato root litter, confirming their importance for litter decomposition. Overall, litter mixing promoted tomato root litter decomposition through altering microbial decomposers, especially through stimulating certain putative keystone taxa. IMPORTANCE Microbial decomposer community plays a key role in litter decomposition, which is an important regulator of soil carbon and nutrient cycling. Though changes in decomposer communities has been proposed as one of the potential underlying mechanisms driving the litter-mixture effects, direct evidence is still lacking. Here, we demonstrated that litter mixing stimulated litter decomposition through altering microbial decomposers at the early stage of decomposition. Moreover, certain putative keystone taxa stimulated by litter mixing contributed to the nonadditive litter-mixture effect. In vitro culturing validated the role of these taxa in litter decomposition. This study also highlights the possibility of regulating litter decomposition through manipulating certain microbial taxa.
Keywords: litter decomposition; litter mixing; litterbags; microbial community; nonadditive effects.
Conflict of interest statement
The authors declare no conflict of interest.
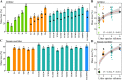
Figures




Similar articles
-
Litter mixing promoted decomposition and altered microbial community in common bean root litter.BMC Microbiol. 2023 May 23;23(1):148. doi: 10.1186/s12866-023-02871-4. BMC Microbiol. 2023. PMID: 37217839 Free PMC article.
-
Temporal dynamics of nutrient release from mulching of legume roots and shoots litter driven by microbial community during decomposition in organic orchards.BMC Plant Biol. 2025 Mar 24;25(1):374. doi: 10.1186/s12870-025-06392-2. BMC Plant Biol. 2025. PMID: 40122813 Free PMC article.
-
Biodiversity mediates the effects of stressors but not nutrients on litter decomposition.Elife. 2020 Jun 26;9:e55659. doi: 10.7554/eLife.55659. Elife. 2020. PMID: 32589139 Free PMC article.
-
[Effects of environmental factors on litter decomposition in arid and semi-arid regions: A review].Ying Yong Sheng Tai Xue Bao. 2013 Nov;24(11):3300-10. Ying Yong Sheng Tai Xue Bao. 2013. PMID: 24564163 Review. Chinese.
-
Functional diversity of terrestrial microbial decomposers and their substrates.C R Biol. 2011 May;334(5-6):393-402. doi: 10.1016/j.crvi.2011.03.001. Epub 2011 May 4. C R Biol. 2011. PMID: 21640948 Review.
Cited by
-
Effect of soil management systems on the rhizosphere bacterial community structure of tobacco: Continuous cropping vs. paddy-upland rotation.Front Plant Sci. 2022 Sep 20;13:996858. doi: 10.3389/fpls.2022.996858. eCollection 2022. Front Plant Sci. 2022. PMID: 36204079 Free PMC article.
-
Microbiome analysis and biocontrol bacteria isolation from rhizosphere soils associated with different sugarcane root rot severity.Front Bioeng Biotechnol. 2022 Dec 16;10:1062351. doi: 10.3389/fbioe.2022.1062351. eCollection 2022. Front Bioeng Biotechnol. 2022. PMID: 36588942 Free PMC article.
-
Litter mixing promoted decomposition and altered microbial community in common bean root litter.BMC Microbiol. 2023 May 23;23(1):148. doi: 10.1186/s12866-023-02871-4. BMC Microbiol. 2023. PMID: 37217839 Free PMC article.
-
Rhizosphere element circling, multifunctionality, aboveground productivity and trade-offs are better predicted by rhizosphere rare taxa.Front Plant Sci. 2022 Sep 8;13:985574. doi: 10.3389/fpls.2022.985574. eCollection 2022. Front Plant Sci. 2022. PMID: 36161026 Free PMC article.
-
Effect of Burkholderia ambifaria LK-P4 inoculation on the plant growth characteristics, metabolism, and pharmacological activity of Anoectochilus roxburghii.Front Plant Sci. 2022 Dec 2;13:1043042. doi: 10.3389/fpls.2022.1043042. eCollection 2022. Front Plant Sci. 2022. PMID: 36531397 Free PMC article.
References
-
- Banerjee S, Kirkby CA, Schmutter D, Bissett A, Kirkegaard JA, Richardson AE. 2016. Network analysis reveals functional redundancy and keystone taxa amongst bacterial and fungal communities during organic matter decomposition in an arable soil. Soil Biol Biochem 97:188–198. doi:10.1016/j.soilbio.2016.03.017. - DOI
-
- Prieto I, Stokes A, Roumet C. 2016. Root functional parameters predict fine root decomposability at the community level. J Ecol 104:725–733. doi:10.1111/1365-2745.12537. - DOI
-
- Pioli S, Sarneel J, Thomas HJD, Domene X, Andres P, Hefting M, Reitz T, Laudon H, Sanden T, Piscova V, Aurela M, Brusetti L. 2020. Linking plant litter microbial diversity to microhabitat conditions, environmental gradients and litter mass loss: Insights from a European study using standard litter bags. Soil Biol Biochem 144:107778. doi:10.1016/j.soilbio.2020.107778. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

