Dcp2 C-terminal cis-binding elements control selective targeting of the decapping enzyme by forming distinct decapping complexes
- PMID: 35604319
- PMCID: PMC9170289
- DOI: 10.7554/eLife.74410
Dcp2 C-terminal cis-binding elements control selective targeting of the decapping enzyme by forming distinct decapping complexes
Abstract
A single Dcp1-Dcp2 decapping enzyme targets diverse classes of yeast mRNAs for decapping-dependent 5' to 3' decay, but the molecular mechanisms controlling mRNA selectivity by the enzyme remain elusive. Through extensive genetic analyses we reveal that Dcp2 C-terminal domain cis-regulatory elements control decapping enzyme target specificity by orchestrating formation of distinct decapping complexes. Two Upf1-binding motifs direct the decapping enzyme to nonsense-mediated mRNA decay substrates, a single Edc3-binding motif targets both Edc3 and Dhh1 substrates, and Pat1-binding leucine-rich motifs target Edc3 and Dhh1 substrates under selective conditions. Although it functions as a unique targeting component of specific complexes, Edc3 is a common component of multiple complexes. Scd6 and Xrn1 also have specific binding sites on Dcp2, allowing them to be directly recruited to decapping complexes. Collectively, our results demonstrate that Upf1, Edc3, Scd6, and Pat1 function as regulatory subunits of the holo-decapping enzyme, controlling both its substrate specificity and enzymatic activation.
Keywords: Dcp2; Edc3; NMD; Pat1; S. cerevisiae; Upf1; Xrn1; chromosomes; gene expression; genetics; genomics.
© 2022, He et al.
Conflict of interest statement
FH, CW No competing interests declared, AJ is co-founder, director, and Scientific Advisory Board chair for PTC Therapeutics Inc
Figures
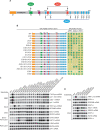




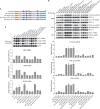


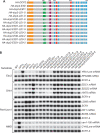
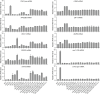






Similar articles
-
Interactions between Upf1 and the decapping factors Edc3 and Pat1 in Saccharomyces cerevisiae.PLoS One. 2011;6(10):e26547. doi: 10.1371/journal.pone.0026547. Epub 2011 Oct 31. PLoS One. 2011. PMID: 22065998 Free PMC article.
-
Control of mRNA decapping by positive and negative regulatory elements in the Dcp2 C-terminal domain.RNA. 2015 Sep;21(9):1633-47. doi: 10.1261/rna.052449.115. Epub 2015 Jul 16. RNA. 2015. PMID: 26184073 Free PMC article.
-
Decapping activators in Saccharomyces cerevisiae act by multiple mechanisms.Mol Cell. 2010 Sep 10;39(5):773-83. doi: 10.1016/j.molcel.2010.08.025. Mol Cell. 2010. PMID: 20832728 Free PMC article.
-
Eukaryotic mRNA decapping factors: molecular mechanisms and activity.FEBS J. 2023 Nov;290(21):5057-5085. doi: 10.1111/febs.16626. Epub 2022 Sep 30. FEBS J. 2023. PMID: 36098474 Free PMC article. Review.
-
mRNA decapping: finding the right structures.Philos Trans R Soc Lond B Biol Sci. 2018 Nov 5;373(1762):20180164. doi: 10.1098/rstb.2018.0164. Philos Trans R Soc Lond B Biol Sci. 2018. PMID: 30397101 Free PMC article. Review.
Cited by
-
Ribosome-bound Upf1 forms distinct 80S complexes and conducts mRNA surveillance.RNA. 2022 Dec;28(12):1621-1642. doi: 10.1261/rna.079416.122. Epub 2022 Oct 3. RNA. 2022. PMID: 36192133 Free PMC article.
-
Human DCP1 is crucial for mRNA decapping and possesses paralog-specific gene regulating functions.Elife. 2024 Nov 1;13:RP94811. doi: 10.7554/eLife.94811. Elife. 2024. PMID: 39485278 Free PMC article.
-
A unique mRNA decapping complex in trypanosomes.Nucleic Acids Res. 2023 Aug 11;51(14):7520-7540. doi: 10.1093/nar/gkad497. Nucleic Acids Res. 2023. PMID: 37309887 Free PMC article.
-
Decapping activators Edc3 and Scd6 act redundantly with Dhh1 in post-transcriptional repression of starvation-induced pathways.bioRxiv [Preprint]. 2024 Aug 28:2024.08.28.610059. doi: 10.1101/2024.08.28.610059. bioRxiv. 2024. PMID: 39257769 Free PMC article. Preprint.
-
RNA anchoring of Upf1 facilitates recruitment of Dcp2 in the NMD decapping complex.Nucleic Acids Res. 2025 Feb 27;53(5):gkaf160. doi: 10.1093/nar/gkaf160. Nucleic Acids Res. 2025. PMID: 40071934 Free PMC article.
References
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

