Distinct patterns of gene expression during regeneration and asexual reproduction in the annelid Pristina leidyi
- PMID: 35604322
- PMCID: PMC9790225
- DOI: 10.1002/jez.b.23143
Distinct patterns of gene expression during regeneration and asexual reproduction in the annelid Pristina leidyi
Abstract
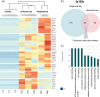
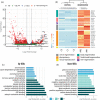
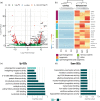
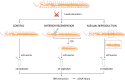
Regeneration, the ability to replace lost body parts, is a widespread phenomenon in the animal kingdom often connected to asexual reproduction or fission, since the only difference between the two appears to be the stimulus that triggers them. Both developmental processes have largely been characterized; however, the molecular toolkit and genetic mechanisms underlying these events remain poorly unexplored. Annelids, in particular the oligochaete Pristina leidyi, provide a good model system to investigate these processes as they show diverse ways to regenerate, and can reproduce asexually through fission under laboratory conditions. Here, we used a comparative transcriptomics approach based on RNA-sequencing and differential gene expression analyses to understand the molecular mechanisms involved in anterior regeneration and asexual reproduction. We found 291 genes upregulated during anterior regeneration, including several regeneration-related genes previously reported in other annelids such as frizzled, paics, and vdra. On the other hand, during asexual reproduction, 130 genes were found upregulated, and unexpectedly, many of them were related to germline development during sexual reproduction. We also found important differences between anterior regeneration and asexual reproduction, with the latter showing a gene expression profile more similar to that of control individuals. Nevertheless, we identified 35 genes that were upregulated in both conditions, many of them related to cell pluripotency, stem cells, and cell proliferation. Overall, our results shed light on the molecular mechanisms that control anterior regeneration and asexual reproduction in annelids and reveal similarities with other animals, suggesting that the genetic machinery controlling these processes is conserved across metazoans.
Keywords: Annelida; Pristina leidyi; differential gene expression; paratomic fission; regeneration; reproduction.
© 2022 The Authors. Journal of Experimental Zoology Part B: Molecular and Developmental Evolution published by Wiley Periodicals LLC.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures

References
-
- Álvarez‐Campos, P. , Kenny, N. J. , Verdes, A. , Fernández, R. , Novo, M. , Giribet, G. , & Riesgo, A. (2019). Delegating sex: Differential gene expression in stolonizing syllids uncovers the hormonal control of reproduction. Genome Biology and Evolution, 11, 295–318. 10.1093/gbe/evy265 - DOI - PMC - PubMed
-
- Andrews, S . (2010). FastQC: Una herramienta de control de calidad para datos de secuencia de alto rendimiento. Accessed February 12, 2021. https://www.bioinformatics.babraham.ac.uk/projects/fastqc
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

