IL-13-programmed airway tuft cells produce PGE2, which promotes CFTR-dependent mucociliary function
- PMID: 35608904
- PMCID: PMC9310525
- DOI: 10.1172/jci.insight.159832
IL-13-programmed airway tuft cells produce PGE2, which promotes CFTR-dependent mucociliary function
Abstract
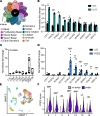
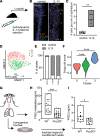
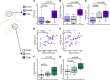
Chronic type 2 (T2) inflammatory diseases of the respiratory tract are characterized by mucus overproduction and disordered mucociliary function, which are largely attributed to the effects of IL-13 on common epithelial cell types (mucus secretory and ciliated cells). The role of rare cells in airway T2 inflammation is less clear, though tuft cells have been shown to be critical in the initiation of T2 immunity in the intestine. Using bulk and single-cell RNA sequencing of airway epithelium and mouse modeling, we found that IL-13 expanded and programmed airway tuft cells toward eicosanoid metabolism and that tuft cell deficiency led to a reduction in airway prostaglandin E2 (PGE2) concentration. Allergic airway epithelia bore a signature of PGE2 activation, and PGE2 activation led to cystic fibrosis transmembrane receptor-dependent ion and fluid secretion and accelerated mucociliary transport. These data reveal a role for tuft cells in regulating epithelial mucociliary function in the allergic airway.
Keywords: Allergy; Chloride channels; Eicosanoids; Inflammation; Pulmonology.
Figures


References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

