The tomato cytochrome P450 CYP712G1 catalyses the double oxidation of orobanchol en route to the rhizosphere signalling strigolactone, solanacol
- PMID: 35612785
- PMCID: PMC9542622
- DOI: 10.1111/nph.18272
The tomato cytochrome P450 CYP712G1 catalyses the double oxidation of orobanchol en route to the rhizosphere signalling strigolactone, solanacol
Abstract
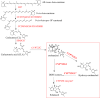
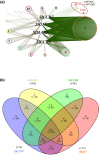
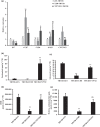
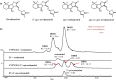
Strigolactones (SLs) are rhizosphere signalling molecules and phytohormones. The biosynthetic pathway of SLs in tomato has been partially elucidated, but the structural diversity in tomato SLs predicts that additional biosynthetic steps are required. Here, root RNA-seq data and co-expression analysis were used for SL biosynthetic gene discovery. This strategy resulted in a candidate gene list containing several cytochrome P450s. Heterologous expression in Nicotiana benthamiana and yeast showed that one of these, CYP712G1, can catalyse the double oxidation of orobanchol, resulting in the formation of three didehydro-orobanchol (DDH) isomers. Virus-induced gene silencing and heterologous expression in yeast showed that one of these DDH isomers is converted to solanacol, one of the most abundant SLs in tomato root exudate. Protein modelling and substrate docking analysis suggest that hydroxy-orbanchol is the likely intermediate in the conversion from orobanchol to the DDH isomers. Phylogenetic analysis demonstrated the occurrence of CYP712G1 homologues in the Eudicots only, which fits with the reports on DDH isomers in that clade. Protein modelling and orobanchol docking of the putative tobacco CYP712G1 homologue suggest that it can convert orobanchol to similar DDH isomers as tomato.
Keywords: CYP712G1; orobanchol; oxidation; solanacol; strigolactone biosynthesis; tomato (Solanum lycopersicum).
© 2022 The Authors. New Phytologist © 2022 New Phytologist Foundation.
Figures



Similar articles
-
The tomato MAX1 homolog, SlMAX1, is involved in the biosynthesis of tomato strigolactones from carlactone.New Phytol. 2018 Jul;219(1):297-309. doi: 10.1111/nph.15131. Epub 2018 Apr 14. New Phytol. 2018. PMID: 29655242
-
Direct conversion of carlactonoic acid to orobanchol by cytochrome P450 CYP722C in strigolactone biosynthesis.Sci Adv. 2019 Dec 18;5(12):eaax9067. doi: 10.1126/sciadv.aax9067. eCollection 2019 Dec. Sci Adv. 2019. PMID: 32064317 Free PMC article.
-
Identification of novel canonical strigolactones produced by tomato.Front Plant Sci. 2022 Dec 14;13:1064378. doi: 10.3389/fpls.2022.1064378. eCollection 2022. Front Plant Sci. 2022. PMID: 36589093 Free PMC article.
-
Stereospecificity in strigolactone biosynthesis and perception.Planta. 2016 Jun;243(6):1361-73. doi: 10.1007/s00425-016-2523-5. Epub 2016 Apr 22. Planta. 2016. PMID: 27105887 Review.
-
Structure Elucidation and Biosynthesis of Orobanchol.Front Plant Sci. 2022 Feb 9;13:835160. doi: 10.3389/fpls.2022.835160. eCollection 2022. Front Plant Sci. 2022. PMID: 35222492 Free PMC article. Review.
Cited by
-
Strigolactones as a hormonal hub for the acclimation and priming to environmental stress in plants.Plant Cell Environ. 2022 Dec;45(12):3611-3630. doi: 10.1111/pce.14461. Epub 2022 Oct 21. Plant Cell Environ. 2022. PMID: 36207810 Free PMC article. Review.
-
OsCYP706C2 diverts rice strigolactone biosynthesis to a noncanonical pathway branch.Sci Adv. 2024 Aug 30;10(35):eadq3942. doi: 10.1126/sciadv.adq3942. Epub 2024 Aug 28. Sci Adv. 2024. PMID: 39196928 Free PMC article.
-
Genome-wide identification of MAXs genes for strigolactones synthesis/signaling in solanaceous plants and analysis of their potential functions in tobacco.PeerJ. 2023 Jan 12;11:e14669. doi: 10.7717/peerj.14669. eCollection 2023. PeerJ. 2023. PMID: 36650839 Free PMC article.
-
Insights into stereoselective ring formation in canonical strigolactone: Identification of a dirigent domain-containing enzyme catalyzing orobanchol synthesis.Proc Natl Acad Sci U S A. 2024 Jun 25;121(26):e2313683121. doi: 10.1073/pnas.2313683121. Epub 2024 Jun 21. Proc Natl Acad Sci U S A. 2024. PMID: 38905237 Free PMC article.
-
Strigolactones and Shoot Branching: What Is the Real Hormone and How Does It Work?Plant Cell Physiol. 2023 Sep 15;64(9):967-983. doi: 10.1093/pcp/pcad088. Plant Cell Physiol. 2023. PMID: 37526426 Free PMC article. Review.
References
-
- Abe S, Sado A, Tanaka K, Kisugi T, Asami K, Ota S, Kim HI, Yoneyama K, Xie X, Ohnishi T et al. 2014. Carlactone is converted to carlactonoic acid by MAX1 in Arabidopsis and its methyl ester can directly interact with AtD14 in vitro . Proceedings of the National Academy of Sciences, USA 111: 18084–18089. - PMC - PubMed
-
- Abuauf H, Haider I, Jia K‐P, Ablazov A, Mi J, Blilou I, Al‐Babili S. 2018. The Arabidopsis DWARF27 gene encodes an all‐trans−/9‐cis‐β‐carotene isomerase and is induced by auxin, abscisic acid and phosphate deficiency. Plant Science 277: 33–42. - PubMed
-
- Akiyama K, Matsuzaki K‐i, Hayashi H. 2005. Plant sesquiterpenes induce hyphal branching in arbuscular mycorrhizal fungi. Nature 435: 824–827. - PubMed
-
- Alder A, Jamil M, Marzorati M, Bruno M, Vermathen M, Bigler P, Ghisla S, Bouwmeester H, Beyer P, Al‐Babili S. 2012. The path from β‐carotene to carlactone, a strigolactone‐like plant hormone. Science 335: 1348–1351. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Research Materials

