Meta-Analysis of Caenorhabditis elegans Transcriptomics Implicates Hedgehog-Like Signaling in Host-Microbe Interactions
- PMID: 35620104
- PMCID: PMC9127769
- DOI: 10.3389/fmicb.2022.853629
Meta-Analysis of Caenorhabditis elegans Transcriptomics Implicates Hedgehog-Like Signaling in Host-Microbe Interactions
Abstract
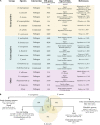
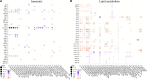
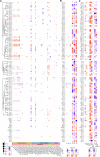
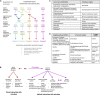
Controlling nematode-caused diseases that affect cattle and crops world-wide remains a critical economic issue, owing to the lack of effective sustainable interventions. The interdependence of roundworms and their environmental microbes, including their microbiota, offers an opportunity for developing more targeted anthelminthic strategies. However, paucity of information and a currently narrow understanding of nematode-microbe interactions limited to specific infection contexts has precluded us from exploiting it. With the advent of omics approaches to map host-microbe genetic interactions, particularly in the model roundworm Caenorhabditis elegans, large datasets are now available across multiple models, that enable identification of nematode-microbe-specific pathways. In this work we collected 20 transcriptomic datasets documenting gene expression changes of C. elegans exposed to 20 different commensal and pathogenic microbes, performing gene enrichment analyses followed by functional testing using RNA interference directed toward genes of interest, before contrasting results from transcriptomic meta-analyses and phenomics. Differential expression analyses revealed a broad enrichment in signaling, innate immune response and (lipid) metabolism genes. Amongst signaling gene families, the nematode-divergent and expanded Hedgehog-like signaling (HHLS) pathway featured prominently. Indeed, 24/60 C. elegans Hedgehog-like proteins (HRPs) and 15/27 Patched-related receptors (PTRs) were differentially expressed in at least four microbial contexts, while up to 32/60 HRPs could be differentially expressed in a single context. interestingly, differentially expressed genes followed a microbe-specific pattern, suggestive of an adaptive microbe-specific response. To investigate this further, we knocked-down 96 individual HHLS genes by RNAi, using high-throughput assays to assess their impact on three worm-gut infection models (Pseudomonas aeruginosa, Staphylococcus aureus, and Enterococcus faecalis) and two worm-commensal paradigms (Comamonas sp., and Bacillus subtilis). We notably identified new putative infection response genes whose upregulation was required for normal pathogen resistance (i.e., grl-21 and ptr-18 protective against E. faecalis), as well as commensal-specific host-gene expression changes that are required for normal host stress handling. Importantly, interactions appeared more microbe-specific than shared. Our results thus implicate the Hedgehog-like signaling pathway in the modulation and possibly fine-tuning of nematode-microbe interactions and support the idea that interventions targeting this pathway may provide a new avenue for anthelmintic development.
Keywords: stress; C. elegans; Hedgehog; LFASS; RNAi; host-microbe interactions; infection; transcriptomics.
Copyright © 2022 Zárate-Potes, Ali, Ribeiro Camacho, Brownless and Benedetto.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Novel Immune Modulators Enhance Caenorhabditis elegans Resistance to Multiple Pathogens.mSphere. 2021 Jan 6;6(1):e00950-20. doi: 10.1128/mSphere.00950-20. mSphere. 2021. PMID: 33408224 Free PMC article.
-
The Caenorhabditis elegans proteome response to two protective Pseudomonas symbionts.mBio. 2024 Apr 10;15(4):e0346323. doi: 10.1128/mbio.03463-23. Epub 2024 Feb 27. mBio. 2024. PMID: 38411078 Free PMC article.
-
Immune-mediated competition benefits protective microbes over pathogens in a novel host species.Heredity (Edinb). 2022 Dec;129(6):327-335. doi: 10.1038/s41437-022-00569-3. Epub 2022 Nov 9. Heredity (Edinb). 2022. PMID: 36352206 Free PMC article.
-
Host-microbe interactions and the behavior of Caenorhabditis elegans.J Neurogenet. 2020 Sep-Dec;34(3-4):500-509. doi: 10.1080/01677063.2020.1802724. Epub 2020 Aug 12. J Neurogenet. 2020. PMID: 32781873 Free PMC article. Review.
-
Genetic and molecular analysis of nematode-microbe interactions.Cell Microbiol. 2011 Apr;13(4):497-507. doi: 10.1111/j.1462-5822.2011.01570.x. Epub 2011 Jan 30. Cell Microbiol. 2011. PMID: 21276170 Review.
Cited by
-
Shigella sonnei and Shigella flexneri infection in Caenorhabditis elegans led to species-specific regulatory responses in the host and pathogen.Microb Genom. 2025 Jan;11(1):001339. doi: 10.1099/mgen.0.001339. Microb Genom. 2025. PMID: 39853209 Free PMC article.
-
C. elegans Hedgehog-related proteins are tissue- and substructure-specific components of the cuticle and pre-cuticle.bioRxiv [Preprint]. 2023 Dec 26:2023.12.26.573316. doi: 10.1101/2023.12.26.573316. bioRxiv. 2023. Update in: Genetics. 2024 Aug 7;227(4):iyae081. doi: 10.1093/genetics/iyae081. PMID: 38234847 Free PMC article. Updated. Preprint.
-
The Caenorhabditis elegans cuticle and precuticle: a model for studying dynamic apical extracellular matrices in vivo.Genetics. 2024 Aug 7;227(4):iyae072. doi: 10.1093/genetics/iyae072. Genetics. 2024. PMID: 38995735 Free PMC article. Review.
-
Caenorhabditis elegans Hedgehog-related proteins are tissue- and substructure-specific components of the cuticle and precuticle.Genetics. 2024 Aug 7;227(4):iyae081. doi: 10.1093/genetics/iyae081. Genetics. 2024. PMID: 38739761 Free PMC article.
-
The longevity response to warm temperature is neurally controlled via the regulation of collagen genes.Aging Cell. 2023 May;22(5):e13815. doi: 10.1111/acel.13815. Epub 2023 Mar 9. Aging Cell. 2023. PMID: 36895142 Free PMC article.
References
LinkOut - more resources
Full Text Sources

