An Insight Based on Computational Analysis of the Interaction between the Receptor-Binding Domain of the Omicron Variants and Human Angiotensin-Converting Enzyme 2
- PMID: 35625526
- PMCID: PMC9171583
- DOI: 10.3390/biology11050797
An Insight Based on Computational Analysis of the Interaction between the Receptor-Binding Domain of the Omicron Variants and Human Angiotensin-Converting Enzyme 2
Abstract
Concerns have been raised about the high number of mutations in the spike protein of the new emergence of the highly transmissible Omicron variant (B.1.1529 lineage) of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). This variant's extraordinary ability to evade antibodies would significantly impair the current vaccination program. This present study aimed to computationally analyze the interaction between the receptor-binding domain (RBD) in the spike protein of Omicron variants and human angiotensin-converting enzyme 2 (hACE2). The docking results indicated that Omicron BA.2 has exceptionally strong interactions with hACE2 in comparison to Omicron BA.1, Delta, and wild-type, as indicated by various parameters such as salt bridge, hydrogen bond, and non-bonded interactions. The results of the molecular dynamics simulation study corroborate these findings, indicating that Omicron BA.2 has a strong and stable interaction with hACE2. This study provides insight into the development of an effective intervention against this variant.
Keywords: Omicron; binding mechanism; computational analysis; receptor-binding domain; spike protein.
Conflict of interest statement
The authors declare no conflict of interest.
Figures






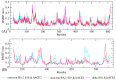
References
-
- Chaguza C., Coppi A., Earnest R., Ferguson D., Kerantzas N., Warner F., Young H.P., Breban M.I., Billig K., Koch R.T., et al. Rapid emergence of SARS-CoV-2 Omicron variant is associated with an infection advantage over Delta in vaccinated persons. medRxiv. 2022:1–21. doi: 10.1016/j.medj.2022.03.010. - DOI - PMC - PubMed
-
- Dejnirattisai W., Huo J., Zhou D., Zahradník J., Supasa P., Liu C., Duyvesteyn H.M.E., Ginn H.M., Mentzer A.J., Tuekprakhon A., et al. Omicron-B.1.1.529 leads to widespread escape from neutralizing antibody responses. bioRxiv Biol. 2021 doi: 10.1101/2021.12.03.471045. preprint . - DOI - PMC - PubMed
-
- VanBlargan L.A., Errico J.M., Halfmann P.J., Zost S.J., Crowe J.E., Purcell L.A., Kawaoka Y., Corti D., Fremont D.H., Diamond M.S. An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by therapeutic monoclonal antibodies. Nat. Med. 2022;28:490–495. doi: 10.1038/s41591-021-01678-y. - DOI - PMC - PubMed
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

