Congenital sideroblastic anemia model due to ALAS2 mutation is susceptible to ferroptosis
- PMID: 35637209
- PMCID: PMC9151922
- DOI: 10.1038/s41598-022-12940-9
Congenital sideroblastic anemia model due to ALAS2 mutation is susceptible to ferroptosis
Abstract
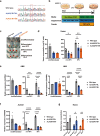
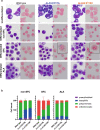
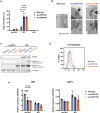
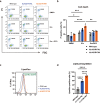
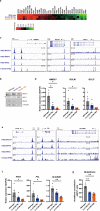
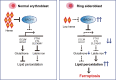
X-linked sideroblastic anemia (XLSA), the most common form of congenital sideroblastic anemia, is caused by a germline mutation in the erythroid-specific 5-aminolevulinate synthase (ALAS2) gene. In XLSA, defective heme biosynthesis leads to ring sideroblast formation because of excess mitochondrial iron accumulation. In this study, we introduced ALAS2 missense mutations on human umbilical cord blood-derived erythroblasts; hereafter, we refer to them as XLSA clones. XLSA clones that differentiated into mature erythroblasts showed an increased frequency of ring sideroblast formation with impaired hemoglobin biosynthesis. The expression profiling revealed significant enrichment of genes involved in ferroptosis, which is a form of regulated cell death induced by iron accumulation and lipid peroxidation. Notably, treatment with erastin, a ferroptosis inducer, caused a higher proportion of cell death in XLSA clones. XLSA clones exhibited significantly higher levels of intracellular lipid peroxides and enhanced expression of BACH1, a regulator of iron metabolism and potential accelerator of ferroptosis. In XLSA clones, BACH1 repressed genes involved in iron metabolism and glutathione synthesis. Collectively, defective heme biosynthesis in XLSA clones could confer enhanced BACH1 expression, leading to increased susceptibility to ferroptosis. The results of our study provide important information for the development of novel therapeutic targets for XLSA.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Generation and Molecular Characterization of Human Ring Sideroblasts: a Key Role of Ferrous Iron in Terminal Erythroid Differentiation and Ring Sideroblast Formation.Mol Cell Biol. 2019 Mar 19;39(7):e00387-18. doi: 10.1128/MCB.00387-18. Print 2019 Apr 1. Mol Cell Biol. 2019. PMID: 30670569 Free PMC article.
-
An erythroid-specific lentiviral vector improves anemia and iron metabolism in a new model of XLSA.Blood. 2025 Jan 2;145(1):98-113. doi: 10.1182/blood.2024025846. Blood. 2025. PMID: 39656107
-
The role of genetic testing in accurate diagnosis of X-linked sideroblastic anemia: novel ALAS2 mutations and the impact of X-chromosome inactivation.Sci Rep. 2025 Apr 7;15(1):11843. doi: 10.1038/s41598-025-95590-x. Sci Rep. 2025. PMID: 40195342 Free PMC article.
-
[Emerging perspectives on sideroblastic anemia].Rinsho Ketsueki. 2025;66(7):557-564. doi: 10.11406/rinketsu.66.557. Rinsho Ketsueki. 2025. PMID: 40769913 Review. Japanese.
-
Molecular pathophysiology and genetic mutations in congenital sideroblastic anemia.Free Radic Biol Med. 2019 Mar;133:179-185. doi: 10.1016/j.freeradbiomed.2018.08.008. Epub 2018 Aug 8. Free Radic Biol Med. 2019. PMID: 30098397 Review.
Cited by
-
Murine models of erythroid 5ALA synthesis disorders and their conditional synthetic lethal dependency on pyridoxine.Blood. 2024 Sep 26;144(13):1418-1432. doi: 10.1182/blood.2023023078. Blood. 2024. PMID: 38900972
-
ATP-Binding Cassette Transporter of Clinical Significance: Sideroblastic Anemia.J Pers Med. 2024 Jun 14;14(6):636. doi: 10.3390/jpm14060636. J Pers Med. 2024. PMID: 38929857 Free PMC article. Review.
-
Metabolomics analysis and mRNA/miRNA profiling reveal potential cardiac regulatory mechanisms in Yili racehorses under different training regimens.PLoS One. 2025 Jul 14;20(7):e0322468. doi: 10.1371/journal.pone.0322468. eCollection 2025. PLoS One. 2025. PMID: 40658689 Free PMC article.
-
The interactions between ineffective erythropoiesis and ferroptosis in β-thalassemia.Front Physiol. 2024 Feb 26;15:1346173. doi: 10.3389/fphys.2024.1346173. eCollection 2024. Front Physiol. 2024. PMID: 38468700 Free PMC article. Review.
-
Ferroptosis regulation by Cap'n'collar family transcription factors.J Biol Chem. 2024 Aug;300(8):107583. doi: 10.1016/j.jbc.2024.107583. Epub 2024 Jul 16. J Biol Chem. 2024. PMID: 39025451 Free PMC article. Review.
References
-
- Bottomley, S. S. & Fleming, M. D. Sideroblastic anemia: diagnosis and management. Hematol. Oncol. Clin. N. Am.28, 653–670, v. 10.1016/j.hoc.2014.04.008 (2014). - PubMed
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

