Platform for isolation and characterization of SARS-CoV-2 variants enables rapid characterization of Omicron in Australia
- PMID: 35637329
- PMCID: PMC9159941
- DOI: 10.1038/s41564-022-01135-7
Platform for isolation and characterization of SARS-CoV-2 variants enables rapid characterization of Omicron in Australia
Abstract
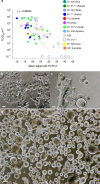
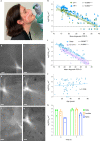
Genetically distinct variants of severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) have emerged since the start of the COVID-19 pandemic. Over this period, we developed a rapid platform (R-20) for viral isolation and characterization using primary remnant diagnostic swabs. This, combined with quarantine testing and genomics surveillance, enabled the rapid isolation and characterization of all major SARS-CoV-2 variants circulating in Australia in 2021. Our platform facilitated viral variant isolation, rapid resolution of variant fitness using nasopharyngeal swabs and ranking of evasion of neutralizing antibodies. In late 2021, variant of concern Omicron (B1.1.529) emerged. Using our platform, we detected and characterized SARS-CoV-2 VOC Omicron. We show that Omicron effectively evades neutralization antibodies and has a different entry route that is TMPRSS2-independent. Our low-cost platform is available to all and can detect all variants of SARS-CoV-2 studied so far, with the main limitation being that our platform still requires appropriate biocontainment.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures




References
-
- Bennett, C. M. Learning to live with COVID-19 in Australia: time for a new approach. Public Health Res. Pract. 10.17061/phrp3132110 (2021). - PubMed
Publication types
MeSH terms
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

