The Dihydrouridine landscape from tRNA to mRNA: a perspective on synthesis, structural impact and function
- PMID: 35638108
- PMCID: PMC9176250
- DOI: 10.1080/15476286.2022.2078094
The Dihydrouridine landscape from tRNA to mRNA: a perspective on synthesis, structural impact and function
Abstract
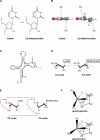
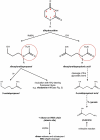
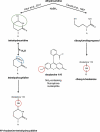
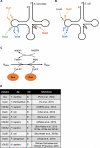
The universal dihydrouridine (D) epitranscriptomic mark results from a reduction of uridine by the Dus family of NADPH-dependent reductases and is typically found within the eponym D-loop of tRNAs. Despite its apparent simplicity, D is structurally unique, with the potential to deeply affect the RNA backbone and many, if not all, RNA-connected processes. The first landscape of its occupancy within the tRNAome was reported 20 years ago. Its potential biological significance was highlighted by observations ranging from a strong bias in its ecological distribution to the predictive nature of Dus enzymes overexpression for worse cancer patient outcomes. The exquisite specificity of the Dus enzymes revealed by a structure-function analyses and accumulating clues that the D distribution may expand beyond tRNAs recently led to the development of new high-resolution mapping methods, including Rho-seq that established the presence of D within mRNAs and led to the demonstration of its critical physiological relevance.
Keywords: Dihydrouridine; Dus; RNA modification; epitranscriptome.
Conflict of interest statement
O.F. is a consultant for Quantoom Biosciences.
Figures

References
-
- Grosjean H. Nucleic acids are not boring long polymers of only four types of nucleotides: a guided tour. DNA and RNA modification enzymes: structure, mechanism, function and evolution. Landes Bioscience. 2009: 1–18.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
