Catchet-MS identifies IKZF1-targeting thalidomide analogues as novel HIV-1 latency reversal agents
- PMID: 35640596
- PMCID: PMC9177988
- DOI: 10.1093/nar/gkac407
Catchet-MS identifies IKZF1-targeting thalidomide analogues as novel HIV-1 latency reversal agents
Abstract
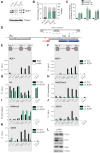
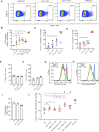
A major pharmacological strategy toward HIV cure aims to reverse latency in infected cells as a first step leading to their elimination. While the unbiased identification of molecular targets physically associated with the latent HIV-1 provirus would be highly valuable to unravel the molecular determinants of HIV-1 transcriptional repression and latency reversal, due to technical limitations, this has been challenging. Here we use a dCas9 targeted chromatin and histone enrichment strategy coupled to mass spectrometry (Catchet-MS) to probe the differential protein composition of the latent and activated HIV-1 5'LTR. Catchet-MS identified known and novel latent 5'LTR-associated host factors. Among these, IKZF1 is a novel HIV-1 transcriptional repressor, required for Polycomb Repressive Complex 2 recruitment to the LTR. We find the clinically advanced thalidomide analogue iberdomide, and the FDA approved analogues lenalidomide and pomalidomide, to be novel LRAs. We demonstrate that, by targeting IKZF1 for degradation, these compounds reverse HIV-1 latency in CD4+ T-cells isolated from virally suppressed people living with HIV-1 and that they are able to synergize with other known LRAs.
© The Author(s) 2022. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



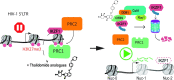
References
-
- Siliciano J.M., Siliciano R.F.. The remarkable stability of the latent reservoir for HIV-1 in resting memory CD4+ T cells. J. Infect. Dis. 2015; 212:1345–1347. - PubMed
-
- Deeks S.G. HIV: shock and kill. Nature. 2012; 487:439–440. - PubMed
-
- Ne E., Palstra R.J., Mahmoudi T.. Transcription: insights from the HIV-1 promoter. Int. Rev. Cell Mol. Biol. 2018; 335:191–243. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

