Automated 3D reconstruction of the fetal thorax in the standard atlas space from motion-corrupted MRI stacks for 21-36 weeks GA range
- PMID: 35649314
- PMCID: PMC7614011
- DOI: 10.1016/j.media.2022.102484
Automated 3D reconstruction of the fetal thorax in the standard atlas space from motion-corrupted MRI stacks for 21-36 weeks GA range
Abstract
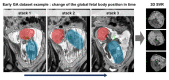
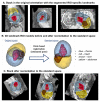
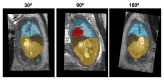
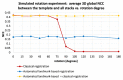
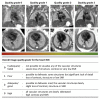
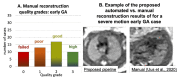
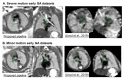
Slice-to-volume registration (SVR) methods allow reconstruction of high-resolution 3D images from multiple motion-corrupted stacks. SVR-based pipelines have been increasingly used for motion correction for T2-weighted structural fetal MRI since they allow more informed and detailed diagnosis of brain and body anomalies including congenital heart defects (Lloyd et al., 2019). Recently, fully automated rigid SVR reconstruction of the fetal brain in the atlas space was achieved in Salehi et al. (2019) that used convolutional neural networks (CNNs) for segmentation and pose estimation. However, these CNN-based methods have not yet been applied to the fetal trunk region. Meanwhile, the existing rigid and deformable SVR (DSVR) solutions (Uus et al., 2020) for the fetal trunk region are limited by the requirement of manual input as well the narrow capture range of the classical gradient descent based registration methods that cannot resolve severe fetal motion frequently occurring at the early gestational age (GA). Furthermore, in our experience, the conventional 2D slice-wise CNN-based brain masking solutions are reportedly prone to errors that require manual corrections when applied on a wide range of acquisition protocols or abnormal cases in clinical setting. In this work, we propose a fully automated pipeline for reconstruction of the fetal thorax region for 21-36 weeks GA range T2-weighted MRI datasets. It includes 3D CNN-based intra-uterine localisation of the fetal trunk and landmark-guided pose estimation steps that allow automated DSVR reconstruction in the standard radiological space irrespective of the fetal trunk position or the regional stack coverage. The additional step for generation of the common template space and rejection of outliers provides the means for automated exclusion of stacks affected by low image quality or extreme motion. The pipeline was quantitatively evaluated on a series of experiments including fetal MRI datasets and simulated rotation motion. Furthermore, we performed a qualitative assessment of the image reconstruction quality in terms of the definition of vascular structures on 100 early (median 23.14 weeks) and late (median 31.79 weeks) GA group MRI datasets covering 21 to 36 weeks GA range.
Keywords: Automated localisation; Automated pose estimation; Deformable slice-to-volume registration; Fetal MRI; Fetal heart.
Copyright © 2022 Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of Competing Interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures












References
-
- Çiçek O, Abdulkadir A, Lienkamp SS, Brox T, Ronneberger O. 3d u-net: learning dense volumetric segmentation from sparse annotation; MICCAI 2016; 2016. pp. 424–432. - DOI
-
- Cordero-Grande L, 1, Price AN, Hughes EJ, Wright R, Rutherford MA, Hajnal JV. Automating motion compensation in 3T fetal brain imaging: localize, align and reconstruct; ISMRM 2019; 2019. p. 1000.
-
- Ebner M, Wang G, Li W, Aertsen M, Patel PA, Aughwane R, Melbourne A, Doel T, Dymarkowski S, De Coppi P, David AL, et al. An automated framework for localization, segmentation and super-resolution reconstruction of fetal brain MRI. Neuroimage. 2020;206:116324. doi: 10.1016/j.neuroimage.2019.116324. - DOI - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

