A mice resting-state functional magnetic resonance imaging dataset on the effects of medetomidine dosages and prior-stimulation on functional connectivity
- PMID: 35651667
- PMCID: PMC9149017
- DOI: 10.1016/j.dib.2022.108279
A mice resting-state functional magnetic resonance imaging dataset on the effects of medetomidine dosages and prior-stimulation on functional connectivity
Abstract
Nine 8 C57Bl6 mice (9 ± 0.5 months) were utilised for this dataset. Each animal was scanned twice on a 9.4T Bruker Magnetic Resonance Imaging (MRI) scanner using a cryogenically cooled coil with 0.1 mg/kg body weight/h (low) or 0.3 mg/kg body weight/h (high) medetomidine doses; 0.5% isoflurane was used in conjunction with both doses. The scans were one week apart, and the first session's dose was decided randomly. In each session, the animal had a pre-stimulation resting-state functional Magnetic Resonance Imaging (rs-fMRI) scan followed by 10 min where mild, constant electrical stimulation to the forepaw was applied, and a post-stimulation rs-fMRI scan. Each fMRI scan lasted 10 min, and there was 5 min break between fMRI scans. The dataset included, for each animal, a pair of forward-phase and reverse-phase gradient echo Echo-Planar-Imaging (EPI) images for EPI distortion correction purpose and three (unprocessed) functional MRI images acquired using the same EPI sequence: prior, during, and post-stimulation. The MRI data was saved in compressed NIFTI format converted from Bruker DICOMs. The dataset also included the pre-processed functional MRI images, with the following pre-processing steps: slice-timing correction, temporal despiking, motion correction, distortion correction, band-pass filtration at 0.01-0.2 Hz, and spatial normalisation. This dataset adds to the publicly available collection of resting-state functional MRI in the mice and facilitates reproducibility and validation of functional imaging and its analysis.
Keywords: Electrical stimulation; Functional connectivity; MRI; Magnetic Resonance Imaging; Medetomidine; Resting state.
© 2022 The Authors. Published by Elsevier Inc.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

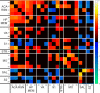
References
-
- Grandjean J., Canella C., Anckaerts C., Ayrancı G., Bougacha S., Bienert T., Buehlmann D., Coletta L., Gallino D., Gass N., Garin C.M., Nadkarni N.A., Hübner N.S., Karatas M., Komaki Y., Kreitz S., Mandino F., Mechling A.E., Sato C., Sauer K., Shah D., Strobelt S., Takata N., Wank I., Wu T., Yahata N., Yeow L.Y., Yee Y., Aoki I., Chakravarty M.M., Chang W.T., Dhenain M., von Elverfeldt D., Harsan L.A., Hess A., Jiang T., Keliris G.A., Lerch J.P., Meyer-Lindenberg A., Okano H., Rudin M., Sartorius A., Van der Linden A., Verhoye M., Weber-Fahr W., Wenderoth N., Zerbi V., Gozzi A. Common functional networks in the mouse brain revealed by multi-centre resting-state fMRI analysis. Neuroimage. 2020:205. doi: 10.1016/j.neuroimage.2019.116278. - DOI - PMC - PubMed
-
- Smith S.M., Jenkinson M., Woolrich M.W., Beckmann C.F., Behrens T.E.J., Johansen-Berg H., Bannister P.R., De Luca M., Drobnjak I., Flitney D.E., Niazy R.K., Saunders J., Vickers J., Zhang Y., De Stefano N., Brady J.M., Matthews P.M. Advances in functional and structural MR image analysis and implementation as FSL. Neuroimage. 2004;23:S208–S219. doi: 10.1016/j.neuroimage.2004.07.051. - DOI - PubMed
Associated data
LinkOut - more resources
Full Text Sources

