Covariation of Pluripotency Markers and Biomechanical Properties in Mouse Embryonic Stem Cells
- PMID: 35652102
- PMCID: PMC9149596
- DOI: 10.3389/fcell.2022.858884
Covariation of Pluripotency Markers and Biomechanical Properties in Mouse Embryonic Stem Cells
Abstract
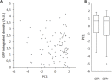
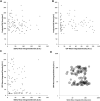
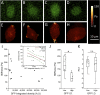
Pluripotent cells are subject to much interest as a source of differentiated cellular material for research models, regenerative medical therapies and novel applications such as lab-cultured meat. Greater understanding of the pluripotent state and control over its differentiation is therefore desirable. The role of biomechanical properties in directing cell fate and cell behavior has been increasingly well described in recent years. However, many of the mechanisms which control cell morphology and mechanical properties in somatic cells are absent from pluripotent cells. We leveraged naturally occurring variation in biomechanical properties and expression of pluripotency genes in murine ESCs to investigate the relationship between these parameters. We observed considerable variation in a Rex1-GFP expression reporter line and found that this variation showed no apparent correlation to cell spreading morphology as determined by circularity, Feret ratio, phase contrast brightness or cell spread area, either on a parameter-by-parameter basis, or when evaluated using a combined metric derived by principal component analysis from the four individual criteria. We further confirmed that cell volume does not co-vary with Rex1-GFP expression. Interestingly, we did find that a subpopulation of cells that were readily detached by gentle agitation collectively exhibited higher expression of Nanog, and reduced LmnA expression, suggesting that elevated pluripotency gene expression may correlate with reduced adhesion to the substrate. Furthermore, atomic force microscopy and quantitative fluorescent imaging revealed a connection between cell stiffness and Rex1-GFP reporter expression. Cells expressing high levels of Rex1-GFP are consistently of a relatively low stiffness, while cells with low levels of Rex1-GFP tend toward higher stiffness values. These observations indicate some interaction between pluripotency gene expression and biomechanical properties, but also support a strong role for other interactions between the cell culture regime and cellular biomechanical properties, occurring independently of the core transcriptional network that supports pluripotency.
Keywords: cell adhesion; cell morphology; cell stiffness; mouse embryonic stem cells; pluripotency marker.
Copyright © 2022 Brookes, Thorpe, Rigby Evans, Keeling and Lee.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures



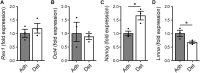
References
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

