Glycogen-autophagy: Molecular machinery and cellular mechanisms of glycophagy
- PMID: 35654138
- PMCID: PMC9249846
- DOI: 10.1016/j.jbc.2022.102093
Glycogen-autophagy: Molecular machinery and cellular mechanisms of glycophagy
Abstract
Autophagy is an essential cellular process involving degradation of superfluous or defective macromolecules and organelles as a form of homeostatic recycling. Initially proposed to be a "bulk" degradation pathway, a more nuanced appreciation of selective autophagy pathways has developed in the literature in recent years. As a glycogen-selective autophagy process, "glycophagy" is emerging as a key metabolic route of transport and delivery of glycolytic fuel substrate. Study of glycophagy is at an early stage. Enhanced understanding of this major noncanonical pathway of glycogen flux will provide important opportunities for new insights into cellular energy metabolism. In addition, glycogen metabolic mishandling is centrally involved in the pathophysiology of several metabolic diseases in a wide range of tissues, including the liver, skeletal muscle, cardiac muscle, and brain. Thus, advances in this exciting new field are of broad multidisciplinary interest relevant to many cell types and metabolic states. Here, we review the current evidence of glycophagy involvement in homeostatic cellular metabolic processes and of molecular mediators participating in glycophagy flux. We integrate information from a variety of settings including cell lines, primary cell culture systems, ex vivo tissue preparations, genetic disease models, and clinical glycogen disease states.
Keywords: Atg8; Gabarapl1; Stbd1; autophagy; glycogen; glycophagy; lysosome.
Copyright © 2022 The Authors. Published by Elsevier Inc. All rights reserved.
Conflict of interest statement
Conflict of interest The authors declare that they have no conflicts of interest with the contents of this article.
Figures

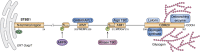
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

