Addressing delayed case reporting in infectious disease forecast modeling
- PMID: 35658007
- PMCID: PMC9200328
- DOI: 10.1371/journal.pcbi.1010115
Addressing delayed case reporting in infectious disease forecast modeling
Abstract
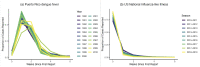
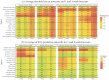
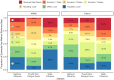
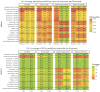
Infectious disease forecasting is of great interest to the public health community and policymakers, since forecasts can provide insight into disease dynamics in the near future and inform interventions. Due to delays in case reporting, however, forecasting models may often underestimate the current and future disease burden. In this paper, we propose a general framework for addressing reporting delay in disease forecasting efforts with the goal of improving forecasts. We propose strategies for leveraging either historical data on case reporting or external internet-based data to estimate the amount of reporting error. We then describe several approaches for adapting general forecasting pipelines to account for under- or over-reporting of cases. We apply these methods to address reporting delay in data on dengue fever cases in Puerto Rico from 1990 to 2009 and to reports of influenza-like illness (ILI) in the United States between 2010 and 2019. Through a simulation study, we compare method performance and evaluate robustness to assumption violations. Our results show that forecasting accuracy and prediction coverage almost always increase when correction methods are implemented to address reporting delay. Some of these methods required knowledge about the reporting error or high quality external data, which may not always be available. Provided alternatives include excluding recently-reported data and performing sensitivity analysis. This work provides intuition and guidance for handling delay in disease case reporting and may serve as a useful resource to inform practical infectious disease forecasting efforts.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures

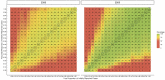
References
-
- Abbott S, Hellewell J, Thompson RN, Sherratt K, Gibbs HP, Bosse NI, et al.. Estimating the time-varying reproduction number of SARS-CoV-2 using national and subnational case counts. Wellcome Open Research. 2020;5:112. doi: 10.12688/wellcomeopenres.16006.2 - DOI
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

