Candidate Phyla Radiation, an Underappreciated Division of the Human Microbiome, and Its Impact on Health and Disease
- PMID: 35658516
- PMCID: PMC9491188
- DOI: 10.1128/cmr.00140-21
Candidate Phyla Radiation, an Underappreciated Division of the Human Microbiome, and Its Impact on Health and Disease
Abstract
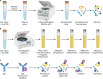
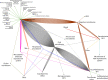
Candidate phyla radiation (CPR) is an emerging division of the bacterial domain within the human microbiota. Still poorly known, these microorganisms were first described in the environment in 1981 as "ultramicrobacteria" with a cell volume under 0.1 μm3 and were first associated with the human oral microbiota in 2007. The evolution of technology has been paramount for the study of CPR within the human microbiota. In fact, since these ultramicrobacteria have yet to be axenically cultured despite ongoing efforts, progress in imaging technology has allowed their observation and morphological description. Although their genomic abilities and taxonomy are still being studied, great strides have been made regarding their taxonomic classification, as well as their lifestyle. In addition, advancements in next-generation sequencing and the continued development of bioinformatics tools have allowed their detection as commensals in different human habitats, including the oral cavity and gastrointestinal and genital tracts, thus highlighting CPR as a nonnegligible part of the human microbiota with an impact on physiological settings. Conversely, several pathologies present dysbiosis affecting CPR levels, including inflammatory, mucosal, and infectious diseases. In this exhaustive review of the literature, we provide a historical perspective on the study of CPR, an overview of the methods available to study these organisms and a description of their taxonomy and lifestyle. In addition, their distribution in the human microbiome is presented in both homeostatic and dysbiotic settings. Future efforts should focus on developing cocultures and, if possible, axenic cultures to obtain isolates and therefore genomes that would provide a better understanding of these ultramicrobacteria, the importance of which in the human microbiome is undeniable.
Keywords: candidate phyla radiation; dysbiosis; genome analysis; human microbiome; imaging; taxonomy.
Conflict of interest statement
The authors declare a conflict of interest. D.R. was a consultant in microbiology for the Hitachi High-Tech Corporation until March 2021. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
Similar articles
-
Ecology of the Oral Microbiome: Beyond Bacteria.Trends Microbiol. 2017 May;25(5):362-374. doi: 10.1016/j.tim.2016.12.012. Epub 2017 Jan 11. Trends Microbiol. 2017. PMID: 28089325 Free PMC article. Review.
-
Structure and Composition of Candidate Phyla Radiation in Supragingival Plaque of Caries Patients.Chin J Dent Res. 2022 Jun 10;25(2):107-118. doi: 10.3290/j.cjdr.b3086339. Chin J Dent Res. 2022. PMID: 35686590
-
State of the Art in the Culture of the Human Microbiota: New Interests and Strategies.Clin Microbiol Rev. 2020 Oct 28;34(1):e00129-19. doi: 10.1128/CMR.00129-19. Print 2020 Dec 16. Clin Microbiol Rev. 2020. PMID: 33115723 Free PMC article. Review.
-
The Roles of Inflammation, Nutrient Availability and the Commensal Microbiota in Enteric Pathogen Infection.Microbiol Spectr. 2015 Jun;3(3). doi: 10.1128/microbiolspec.MBP-0008-2014. Microbiol Spectr. 2015. PMID: 26185088
-
Chemotherapy-induced oral mucositis is associated with detrimental bacterial dysbiosis.Microbiome. 2019 Apr 25;7(1):66. doi: 10.1186/s40168-019-0679-5. Microbiome. 2019. PMID: 31018870 Free PMC article.
Cited by
-
Microscopic and metatranscriptomic analyses revealed unique cross-domain parasitism between phylum Candidatus Patescibacteria/candidate phyla radiation and methanogenic archaea in anaerobic ecosystems.mBio. 2024 Mar 13;15(3):e0310223. doi: 10.1128/mbio.03102-23. Epub 2024 Feb 7. mBio. 2024. PMID: 38323857 Free PMC article.
-
Preliminary landscape of Candidatus Saccharibacteria in the human microbiome.Front Cell Infect Microbiol. 2023 Jul 27;13:1195679. doi: 10.3389/fcimb.2023.1195679. eCollection 2023. Front Cell Infect Microbiol. 2023. PMID: 37577371 Free PMC article.
-
Episymbiotic Saccharibacteria suppresses epithelial immunoactivation through Type IV pili and TLR2 dependent endocytosis.bioRxiv [Preprint]. 2025 Jun 2:2025.05.30.656655. doi: 10.1101/2025.05.30.656655. bioRxiv. 2025. PMID: 40501963 Free PMC article. Preprint.
-
Microbial Symphony: Navigating the Intricacies of the Human Oral Microbiome and Its Impact on Health.Microorganisms. 2024 Mar 13;12(3):571. doi: 10.3390/microorganisms12030571. Microorganisms. 2024. PMID: 38543622 Free PMC article. Review.
-
Alteration of taste perception, food neophobia and oral microbiota composition in children with food allergy.Sci Rep. 2023 Apr 28;13(1):7010. doi: 10.1038/s41598-023-34113-y. Sci Rep. 2023. PMID: 37117251 Free PMC article.
References
-
- Rheims H, Rainey FA, Stackebrandt E. 1996. A molecular approach to search for diversity among bacteria in the environment. J Indust Microbiol Biotechnol 17:159–169. 10.1007/BF01574689. - DOI
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

