Stress-Induced Mutagenesis, Gambler Cells, and Stealth Targeting Antibiotic-Induced Evolution
- PMID: 35658528
- PMCID: PMC9239211
- DOI: 10.1128/mbio.01074-22
Stress-Induced Mutagenesis, Gambler Cells, and Stealth Targeting Antibiotic-Induced Evolution
Abstract
Mechanisms of evolution and evolution of antibiotic resistance are both fundamental and world health problems. Stress-induced mutagenesis defines mechanisms of mutagenesis upregulated by stress responses, which drive adaptation when cells are maladapted to their environments-when stressed. Work in mutagenesis induced by antibiotics had produced tantalizing clues but not coherent mechanisms. We review recent advances in antibiotic-induced mutagenesis that integrate how reactive oxygen species (ROS), the SOS and general stress responses, and multichromosome cells orchestrate a stress response-induced switch from high-fidelity to mutagenic repair of DNA breaks. Moreover, while sibling cells stay stable, a mutable "gambler" cell subpopulation is induced by differentially generated ROS, which signal the general stress response. We discuss other evolvable subpopulations and consider diverse evolution-promoting molecules as potential targets for drugs to slow evolution of antibiotic resistance, cross-resistance, and immune evasion. An FDA-approved drug exemplifies "stealth" evolution-slowing drugs that avoid selecting resistance to themselves or antibiotics.
Keywords: antibiotic resistance; antibiotics; antievolvability drugs; cell subpopulations; drug resistance evolution; evolution; evolvability; stress-induced mutagenesis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures
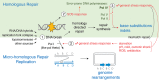

Similar articles
-
Gamblers: An Antibiotic-Induced Evolvable Cell Subpopulation Differentiated by Reactive-Oxygen-Induced General Stress Response.Mol Cell. 2019 May 16;74(4):785-800.e7. doi: 10.1016/j.molcel.2019.02.037. Epub 2019 Apr 1. Mol Cell. 2019. PMID: 30948267 Free PMC article.
-
ppGpp and RNA-polymerase backtracking guide antibiotic-induced mutable gambler cells.Mol Cell. 2023 Apr 20;83(8):1298-1310.e4. doi: 10.1016/j.molcel.2023.03.003. Epub 2023 Mar 24. Mol Cell. 2023. PMID: 36965481 Free PMC article.
-
Persistent damaged bases in DNA allow mutagenic break repair in Escherichia coli.PLoS Genet. 2017 Jul 20;13(7):e1006733. doi: 10.1371/journal.pgen.1006733. eCollection 2017 Jul. PLoS Genet. 2017. PMID: 28727736 Free PMC article.
-
Targeting evolution to inhibit antibiotic resistance.FEBS J. 2020 Oct;287(20):4341-4353. doi: 10.1111/febs.15370. Epub 2020 Jun 8. FEBS J. 2020. PMID: 32434280 Free PMC article. Review.
-
Mutation as a stress response and the regulation of evolvability.Crit Rev Biochem Mol Biol. 2007 Sep-Oct;42(5):399-435. doi: 10.1080/10409230701648502. Crit Rev Biochem Mol Biol. 2007. PMID: 17917874 Free PMC article. Review.
Cited by
-
Metabolic mutations reduce antibiotic susceptibility of E. coli by pathway-specific bottlenecks.Mol Syst Biol. 2025 Mar;21(3):274-293. doi: 10.1038/s44320-024-00084-z. Epub 2025 Jan 2. Mol Syst Biol. 2025. PMID: 39748127 Free PMC article.
-
The global RNA-RNA interactome of Klebsiella pneumoniae unveils a small RNA regulator of cell division.Proc Natl Acad Sci U S A. 2024 Feb 27;121(9):e2317322121. doi: 10.1073/pnas.2317322121. Epub 2024 Feb 20. Proc Natl Acad Sci U S A. 2024. PMID: 38377209 Free PMC article.
-
Microfluidic Ecology Unravels the Genetic and Ecological Drivers of T4r Bacteriophage Resistance in E. coli: Insights into Biofilm-Mediated Evolution.Res Sq [Preprint]. 2024 May 24:rs.3.rs-4356333. doi: 10.21203/rs.3.rs-4356333/v1. Res Sq. 2024. PMID: 38826273 Free PMC article. Preprint.
-
Bioenergetic stress potentiates antimicrobial resistance and persistence.Nat Commun. 2025 Jun 9;16(1):5111. doi: 10.1038/s41467-025-60302-6. Nat Commun. 2025. PMID: 40490453 Free PMC article.
-
Impact of low-intensity 463 nm blue light on proliferation and adaptive mutation of Escherichia coli DH5α cells.Sci Rep. 2025 Jul 1;15(1):20848. doi: 10.1038/s41598-025-06596-4. Sci Rep. 2025. PMID: 40594907 Free PMC article.
References
-
- Murray CJL, Ikuta KS, Sharara F, Swetschinski L, Robles Aguilar G, Gray A, Han C, Bisignano C, Rao P, Wool E, Johnson SC, Browne AJ, Chipeta MG, Fell F, Hackett S, Haines-Woodhouse G, Kashef Hamadani BH, Kumaran EAP, McManigal B, Agarwal R, Akech S, Albertson S, Amuasi J, Andrews J, Aravkin A, Ashley E, Bailey F, Baker S, Basnyat B, Bekker A, Bender R, Bethou A, Bielicki J, Boonkasidecha S, Bukosia J, Carvalheiro C, Castañeda-Orjuela C, Chansamouth V, Chaurasia S, Chiurchiù S, Chowdhury F, Cook AJ, Cooper B, Cressey TR, Criollo-Mora E, Cunningham M, Darboe S, Day NPJ, De Luca M, Dokova K, et al. . 2022. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 399:629–655. doi:10.1016/S0140-6736(21)02724-0. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases

