Peptide vaccine-treated, long-term surviving cancer patients harbor self-renewing tumor-specific CD8+ T cells
- PMID: 35660746
- PMCID: PMC9166698
- DOI: 10.1038/s41467-022-30861-z
Peptide vaccine-treated, long-term surviving cancer patients harbor self-renewing tumor-specific CD8+ T cells
Abstract
The behaviors and fates of immune cells in cancer patients, such as dysfunction and stem-like states leading to memory formation in T cells, are in intense focus of investigation. Here we show, by post hoc analysis of peripheral blood lymphocytes of hepatocellular carcinoma patients previously undergoing vaccination with tumour-associated antigen-derived peptides in our clinical trials (registration numbers UMIN000003511, UMIN000004540, UMIN000005677, UMIN000003514 and UMIN000005678), that induced peptide-specific T cell responses may persist beyond 10 years following vaccination. Tracking TCR clonotypes at the single cell level reveals in two patients that peptide-specific long-lasting CD8+ T cells acquire an effector memory phenotype that associates with cell cycle-related genes (CCNA2 and CDK1), and are characterized by high expression of IL7R, SELL, and NOSIP along with a later stage promotion of the AP-1 transcription factor network (5 years or more past vaccination). We conclude that effective anti-tumor immunity is governed by potentially proliferative memory T cells, specific to cancer antigens.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures



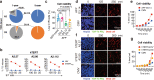

References
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

