Expression and Purification of Bsa XI Restriction Endonuclease and Engineering New Specificity From Bsa XI Specificity Subunit
- PMID: 35663886
- PMCID: PMC9159934
- DOI: 10.3389/fmicb.2022.888435
Expression and Purification of Bsa XI Restriction Endonuclease and Engineering New Specificity From Bsa XI Specificity Subunit
Abstract
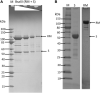
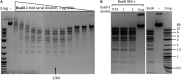
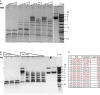
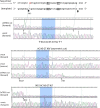
It is stated that BsaXI is a Type IIB restriction endonuclease (REase) that cleaves both sides of its recognition sequence 5'↓N9 AC N5 CTCC N10↓ 3' (complement strand 5' ↓N7 GGAG N5 GT N12↓ 3'), creating 3-base 3' overhangs. Here we report the cloning and expression of bsaXIS and bsaXIRM genes in Escherichia coli. The BsaXI activity was successfully reconstituted by mixing the BsaXI RM fusion subunit with the BsaXI S subunit and the enzyme complex further purified by chromatography over 6 columns. As expected, the S subunit consisted of two subdomains encoding TRD1-CR1 [target recognition domain (TRD), conserved region (CR)] for 5' AC 3', and TRD2-CR2 presumably specifying 5' CTCC 3'. TRD1-CR1 (TRD2-CR2 deletion) or duplication of TRD1 (TRD1-CR1-TRD1-CR2) both generated a new specificity 5' AC N5 GT 3' when the S variants were complexed with the RM subunits. The circular permutation of TRD1 and TRD2, i.e., the relocation of TRD2-CR2 to the N-terminus and TRD1-CR1 to the C-terminus generated the same specificity with the RM subunits, although some wobble cleavage was detected. The TRD2 domain in the BsaXI S subunit can be substituted by a close homolog (∼59% sequence identity) and generated the same specificity. However, TRD2-CR2 domain alone failed to express in E. coli, but CR1-TRD2-CR2 protein could be expressed and purified which showed partial nicking activity with the RM subunits. This work demonstrated that like Type I restriction systems, the S subunit of a Type IIB system could also be manipulated to create new specificities. The genome mining of BsaXI TRD2 homologs in GenBank found more than 36 orphan TRD2 homologs, implying that quite a few orphan TRD2s are present in microbial genomes that may be potentially paired with other TRDs to create new restriction specificities.
Keywords: BsaXI specificity; RM fusion and S subunits; TRD1-CR1; Type IIB restriction endonuclease; circular permutation of BsaXI S subunit.
Copyright © 2022 Gidwani, Heiter and Xu.
Conflict of interest statement
DH and S-YX are employees of New England Biolabs, Inc., a company that develops restriction enzymes and other reagents for research and diagnostic applications. The remaining author declares that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures

Similar articles
-
Cryo-EM structures of DNA-free and DNA-bound BsaXI: architecture of a Type IIB restriction-modification enzyme.Nucleic Acids Res. 2025 Apr 10;53(7):gkaf291. doi: 10.1093/nar/gkaf291. Nucleic Acids Res. 2025. PMID: 40239994 Free PMC article.
-
Domain movement within a gene: a novel evolutionary mechanism for protein diversification.PLoS One. 2011 Apr 14;6(4):e18819. doi: 10.1371/journal.pone.0018819. PLoS One. 2011. PMID: 21533192 Free PMC article.
-
Recognition and discrimination of target mRNAs by Sib RNAs, a cis-encoded sRNA family.Nucleic Acids Res. 2010 Sep;38(17):5851-66. doi: 10.1093/nar/gkq292. Epub 2010 May 7. Nucleic Acids Res. 2010. PMID: 20453032 Free PMC article.
-
Engineering Infrequent DNA Nicking Endonuclease by Fusion of a BamHI Cleavage-Deficient Mutant and a DNA Nicking Domain.Front Microbiol. 2022 Feb 1;12:787073. doi: 10.3389/fmicb.2021.787073. eCollection 2021. Front Microbiol. 2022. PMID: 35178039 Free PMC article.
-
Deficiencies of human C3 complement receptors type 1 (CR1, CD35) and type 2 (CR2, CD21).Immunodefic Rev. 1990;2(1):17-41. Immunodefic Rev. 1990. PMID: 2164822 Review.
Cited by
-
Cryo-EM structures of DNA-free and DNA-bound BsaXI: architecture of a Type IIB restriction-modification enzyme.Nucleic Acids Res. 2025 Apr 10;53(7):gkaf291. doi: 10.1093/nar/gkaf291. Nucleic Acids Res. 2025. PMID: 40239994 Free PMC article.
References
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

