pH sensing controls tissue inflammation by modulating cellular metabolism and endo-lysosomal function of immune cells
- PMID: 35668320
- PMCID: PMC9720675
- DOI: 10.1038/s41590-022-01231-0
pH sensing controls tissue inflammation by modulating cellular metabolism and endo-lysosomal function of immune cells
Abstract
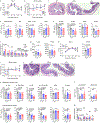
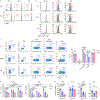
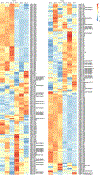
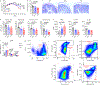
Extracellular acidification occurs in inflamed tissue and the tumor microenvironment; however, a systematic study on how pH sensing contributes to tissue homeostasis is lacking. In the present study, we examine cell type-specific roles of the pH sensor G protein-coupled receptor 65 (GPR65) and its inflammatory disease-associated Ile231Leu-coding variant in inflammation control. GPR65 Ile231Leu knock-in mice are highly susceptible to both bacterial infection-induced and T cell-driven colitis. Mechanistically, GPR65 Ile231Leu elicits a cytokine imbalance through impaired helper type 17 T cell (TH17 cell) and TH22 cell differentiation and interleukin (IL)-22 production in association with altered cellular metabolism controlled through the cAMP-CREB-DGAT1 axis. In dendritic cells, GPR65 Ile231Leu elevates IL-12 and IL-23 release at acidic pH and alters endo-lysosomal fusion and degradation capacity, resulting in enhanced antigen presentation. The present study highlights GPR65 Ile231Leu as a multistep risk factor in intestinal inflammation and illuminates a mechanism by which pH sensing controls inflammatory circuits and tissue homeostasis.
© 2022. The Author(s), under exclusive licence to Springer Nature America, Inc.
Conflict of interest statement
COMPETING INTERESTS
R.J.X. is co-founder of Jnana Therapeutics and Celsius Therapeutics, and M.J.D. is a scientific founder of Maze Therapeutics; these organizations had no roles in this study. The remaining authors declare no competing interests.
Figures











Comment in
-
When inflammation turns sour on T cells.Nat Immunol. 2022 Jul;23(7):991-993. doi: 10.1038/s41590-022-01241-y. Nat Immunol. 2022. PMID: 35697839 No abstract available.
References
-
- Levin LR & Buck J Physiological roles of acid-base sensors. Annu Rev Physiol 77, 347–362 (2015). - PubMed
-
- Ananthakrishnan AN et al. Environmental triggers in IBD: a review of progress and evidence. Nat Rev Gastroenterol Hepatol 15, 39–49 (2018). - PubMed
-
- de Souza HS & Fiocchi C Immunopathogenesis of IBD: current state of the art. Nat Rev Gastroenterol Hepatol 13, 13–27 (2016). - PubMed
METHODS-ONLY REFERENCES
-
- Fleming SJ, Marioni JC & Babadi M CellBender remove-background: a deep generative model for unsupervised removal of background noise from scRNA-seq datasets. bioRxiv 791699 (2019).
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

