Integrating 3D genomic and epigenomic data to enhance target gene discovery and drug repurposing in transcriptome-wide association studies
- PMID: 35672318
- PMCID: PMC9171100
- DOI: 10.1038/s41467-022-30956-7
Integrating 3D genomic and epigenomic data to enhance target gene discovery and drug repurposing in transcriptome-wide association studies
Abstract
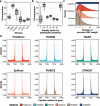
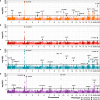
Transcriptome-wide association studies (TWAS) are popular approaches to test for association between imputed gene expression levels and traits of interest. Here, we propose an integrative method PUMICE (Prediction Using Models Informed by Chromatin conformations and Epigenomics) to integrate 3D genomic and epigenomic data with expression quantitative trait loci (eQTL) to more accurately predict gene expressions. PUMICE helps define and prioritize regions that harbor cis-regulatory variants, which outperforms competing methods. We further describe an extension to our method PUMICE +, which jointly combines TWAS results from single- and multi-tissue models. Across 79 traits, PUMICE + identifies 22% more independent novel genes and increases median chi-square statistics values at known loci by 35% compared to the second-best method, as well as achieves the narrowest credible interval size. Lastly, we perform computational drug repurposing and confirm that PUMICE + outperforms other TWAS methods.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures





Similar articles
-
Integrating single cell expression quantitative trait loci summary statistics to understand complex trait risk genes.Nat Commun. 2024 May 20;15(1):4260. doi: 10.1038/s41467-024-48143-1. Nat Commun. 2024. PMID: 38769300 Free PMC article.
-
Influence of tissue context on gene prioritization for predicted transcriptome-wide association studies.Pac Symp Biocomput. 2019;24:296-307. Pac Symp Biocomput. 2019. PMID: 30864331 Free PMC article. Clinical Trial.
-
Leveraging expression from multiple tissues using sparse canonical correlation analysis and aggregate tests improves the power of transcriptome-wide association studies.PLoS Genet. 2021 Apr 8;17(4):e1008973. doi: 10.1371/journal.pgen.1008973. eCollection 2021 Apr. PLoS Genet. 2021. PMID: 33831007 Free PMC article.
-
Opportunities and challenges for transcriptome-wide association studies.Nat Genet. 2019 Apr;51(4):592-599. doi: 10.1038/s41588-019-0385-z. Epub 2019 Mar 29. Nat Genet. 2019. PMID: 30926968 Free PMC article. Review.
-
Integrative Analysis of Multi-omics Data for Discovery and Functional Studies of Complex Human Diseases.Adv Genet. 2016;93:147-90. doi: 10.1016/bs.adgen.2015.11.004. Epub 2016 Jan 25. Adv Genet. 2016. PMID: 26915271 Free PMC article. Review.
Cited by
-
TWAS-GKF: a novel method for causal gene identification in transcriptome-wide association studies with knockoff inference.Bioinformatics. 2024 Aug 2;40(8):btae502. doi: 10.1093/bioinformatics/btae502. Bioinformatics. 2024. PMID: 39189955 Free PMC article.
-
FarmGTEx TWAS-server: An Interactive Web Server for Customized TWAS Analysis.Genomics Proteomics Bioinformatics. 2025 May 10;23(1):qzaf006. doi: 10.1093/gpbjnl/qzaf006. Genomics Proteomics Bioinformatics. 2025. PMID: 39932890 Free PMC article.
-
SUMMIT-FA: a new resource for improved transcriptome imputation using functional annotations.Hum Mol Genet. 2024 Mar 20;33(7):624-635. doi: 10.1093/hmg/ddad205. Hum Mol Genet. 2024. PMID: 38129112 Free PMC article.
-
Integrating single cell expression quantitative trait loci summary statistics to understand complex trait risk genes.Nat Commun. 2024 May 20;15(1):4260. doi: 10.1038/s41467-024-48143-1. Nat Commun. 2024. PMID: 38769300 Free PMC article.
-
Leveraging Random Effects in Cistrome-Wide Association Studies for Decoding the Genetic Determinants of Prostate Cancer.Adv Sci (Weinh). 2024 Sep;11(36):e2400815. doi: 10.1002/advs.202400815. Epub 2024 Aug 5. Adv Sci (Weinh). 2024. PMID: 39099406 Free PMC article.
References
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources

