Biochemical Characterization of SARS-CoV-2 Spike RBD Mutations and Their Impact on ACE2 Receptor Binding
- PMID: 35677879
- PMCID: PMC9168323
- DOI: 10.3389/fmolb.2022.893843
Biochemical Characterization of SARS-CoV-2 Spike RBD Mutations and Their Impact on ACE2 Receptor Binding
Abstract
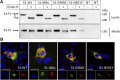
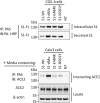
Infection of mammalian cells by SARS-CoV-2 coronavirus requires primary interaction between the receptor binding domain (RBD) of the viral spike protein and the host cell surface receptor angiotensin-converting enzyme 2 (ACE2) glycoprotein. Several mutations in the RBD of SARS-CoV-2 spike protein have been reported for several variants and resulted in wide spread of the COVID pandemic. For instance, the double mutations L452R and E484Q present in the Indian B.1.617 variant have been suggested to cause evasion of the host immune response. The common RBD mutations N501Y and E484K were found to enhance the interaction with the ACE2 receptor. In the current study, we analyzed the biosynthesis and secretion of the RBD double mutants L452R and E484Q in comparison to the wild-type RBD and the individual mutations N501 and E484K in mammalian cells. Moreover, we evaluated the interaction of these variants with ACE2 by means of expression of the S protein and co-immunoprecipitation with ACE2. Our results revealed that the double RBD mutations L452R and E484Q resulted in a higher expression level and secretion of spike S1 protein than other mutations. In addition, an increased interaction of these mutant forms with ACE2 in Calu3 cells was observed. Altogether, our findings highlight the impact of continuous S1 mutations on the pathogenicity of SARS-CoV-2 and provide further biochemical evidence for the dominance and high transmissibility of the double Indian mutations.
Keywords: ACE2 interaction; RBD; SARS-CoV-2; double mutant; spike; transmissibility.
Copyright © 2022 Hoter and Naim.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


References
LinkOut - more resources
Full Text Sources
Miscellaneous

