c-di-AMP Accumulation Regulates Growth, Metabolism, and Immunogenicity of Mycobacterium smegmatis
- PMID: 35685938
- PMCID: PMC9171234
- DOI: 10.3389/fmicb.2022.865045
c-di-AMP Accumulation Regulates Growth, Metabolism, and Immunogenicity of Mycobacterium smegmatis
Abstract
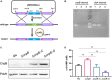
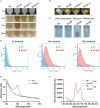
Cyclic dimeric adenosine monophosphate (c-di-AMP) is a ubiquitous second messenger of bacteria involved in diverse physiological processes as well as host immune responses. MSMEG_2630 is a c-di-AMP phosphodiesterase (cnpB) of Mycobacterium smegmatis, which is homologous to Mycobacterium tuberculosis Rv2837c. In this study, cnpB-deleted (ΔcnpB), -complemented (ΔcnpB::C), and -overexpressed (ΔcnpB::O) strains of M. smegmatis were constructed to investigate the role of c-di-AMP in regulating mycobacterial physiology and immunogenicity. This study provides more precise evidence that elevated c-di-AMP level resulted in smaller colonies, shorter bacteria length, impaired growth, and inhibition of potassium transporter in M. smegmatis. This is the first study to report that elevated c-di-AMP level could inhibit biofilm formation and induce porphyrin accumulation in M. smegmatis by regulating associated gene expressions, which may have effects on drug resistance and virulence of mycobacterium. Moreover, the cnpB-deleted strain with an elevated c-di-AMP level could induce enhanced Th1 immune responses after M. tuberculosis infection. Further, the pathological changes and the bacteria burden in ΔcnpB group were comparable with the wild-type M. smegmatis group against M. tuberculosis venous infection in the mouse model. Our findings enhanced the understanding of the physiological role of c-di-AMP in mycobacterium, and M. smegmatis cnpB-deleted strain with elevated c-di-AMP level showed the potential for a vaccine against tuberculosis.
Keywords: M. tuberculosis; Mycobacterium smegmatis; c-di-AMP; immunogenicity; physiology.
Copyright © 2022 Ning, Liang, Xie, Bai, Zhang, Wang, Kang, Lu, Ma, Bai and Bai.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures






Similar articles
-
Cyclic-di-AMP Phosphodiesterase Elicits Protective Immune Responses Against Mycobacterium tuberculosis H37Ra Infection in Mice.Front Cell Infect Microbiol. 2022 Jun 22;12:871135. doi: 10.3389/fcimb.2022.871135. eCollection 2022. Front Cell Infect Microbiol. 2022. PMID: 35811674 Free PMC article.
-
Deletion of the cyclic di-AMP phosphodiesterase gene (cnpB) in Mycobacterium tuberculosis leads to reduced virulence in a mouse model of infection.Mol Microbiol. 2014 Jul;93(1):65-79. doi: 10.1111/mmi.12641. Epub 2014 May 23. Mol Microbiol. 2014. PMID: 24806618 Free PMC article.
-
Cyclic di-AMP-mediated interaction between Mycobacterium tuberculosis ΔcnpB and macrophages implicates a novel strategy for improving BCG vaccination.Pathog Dis. 2018 Mar 1;76(2). doi: 10.1093/femspd/fty008. Pathog Dis. 2018. PMID: 29394352
-
[Advances in Research on the Role of Bacterial c-di-AMP in Host Immunity].Sichuan Da Xue Xue Bao Yi Xue Ban. 2022 Nov;53(6):1098-1103. doi: 10.12182/20220860102. Sichuan Da Xue Xue Bao Yi Xue Ban. 2022. PMID: 36443059 Free PMC article. Review. Chinese.
-
The role of bacterial cyclic di-adenosine monophosphate in the host immune response.Front Microbiol. 2022 Aug 29;13:958133. doi: 10.3389/fmicb.2022.958133. eCollection 2022. Front Microbiol. 2022. PMID: 36106081 Free PMC article. Review.
Cited by
-
Cyclic di-AMP as endogenous adjuvant enhanced BCG-induced trained immunity and protection against Mycobacterium tuberculosis in mice.Front Immunol. 2022 Aug 23;13:943667. doi: 10.3389/fimmu.2022.943667. eCollection 2022. Front Immunol. 2022. PMID: 36081510 Free PMC article.
-
Structures of the DarR transcription regulator reveal unique modes of second messenger and DNA binding.Nat Commun. 2023 Nov 9;14(1):7239. doi: 10.1038/s41467-023-42823-0. Nat Commun. 2023. PMID: 37945601 Free PMC article.
-
Fighting Tuberculosis: In Search of a BCG Replacement.Microorganisms. 2022 Dec 23;11(1):51. doi: 10.3390/microorganisms11010051. Microorganisms. 2022. PMID: 36677343 Free PMC article. Review.
References
-
- Alves Da Silva D., Cavalcanti M. A., Muniz, De Oliveira F., Trentini M. M., Junqueira-Kipnis A. P., et al. (2014). Immunogenicity of a recombinant Mycobacterium smegmatis vaccine expressing the fusion protein CMX in cattle from Goias State. Brazil. J. Vet. Med. Sci. 76 977–984. 10.1292/jvms.13-0338 - DOI - PMC - PubMed
-
- Bai Y., Bai G. (2020). “Cyclic di-AMP in Mycobacterium tuberculosis,” in Microbial Cyclic Di-Nucleotide Signaling, ed. Shan-Ho Chou N. G., Vincent T. (Berlin: Springer; ), 443–454.
LinkOut - more resources
Full Text Sources

