Population level SARS-CoV-2 fecal shedding rates determined via wastewater-based epidemiology
- PMID: 35688254
- PMCID: PMC9172256
- DOI: 10.1016/j.scitotenv.2022.156535
Population level SARS-CoV-2 fecal shedding rates determined via wastewater-based epidemiology
Abstract
Wastewater-based epidemiology (WBE) has been utilized as an early warning tool to anticipate disease outbreaks, especially during the COVID-19 pandemic. However, COVID-19 disease models built from wastewater-collected data have been limited by the complexities involved in estimating SARS-CoV-2 fecal shedding rates. In this study, wastewater from six municipalities in Arizona and Florida with distinct demographics were monitored for SARS-CoV-2 RNA between September 2020 and December 2021. Virus concentrations with corresponding clinical case counts were utilized to estimate community-wide fecal shedding rates that encompassed all infected individuals. Analyses suggest that average SARS-CoV-2 RNA fecal shedding rates typically occurred within a consistent range (7.53-9.29 log10 gc/g-feces); and yet, were unique to each community and influenced by population demographics. Age, ethnicity, and socio-economic factors may have influenced shedding rates. Interestingly, populations with median age between 30 and 39 had the greatest fecal shedding rates. Additionally, rates remained relatively constant throughout the pandemic provided conditions related to vaccination and variants were unchanged. Rates significantly increased in some communities when the Delta variant became predominant. Findings in this study suggest that community-specific shedding rates may be appropriate in model development relating wastewater virus concentrations to clinical case counts.
Keywords: COVID-19; Community demographics; Fecal shedding; SARS-CoV-2; Wastewater-based epidemiology.
Copyright © 2022 Elsevier B.V. All rights reserved.
Conflict of interest statement
Declaration of competing interest The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures

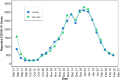

References
-
- Ahmed W., Angel N., Edson J., Bibby K., Bivins A., O’Brien J.W., Choi P.M., Kitajima M., Simpson S.L., Li J., Tscharke B., Verhagen R., Smith W.J.M., Zaugg J., Dierens L., Hugenholtz P., Thomas K.V., Mueller J.F. First confirmed detection of SARS-CoV-2 in untreated wastewater in Australia: a proof of concept for the wastewater surveillance of COVID-19 in the community. Sci. Total Environ. 2020;728 doi: 10.1016/j.scitotenv.2020.138764. - DOI - PMC - PubMed
-
- Ahmed W., Bertsch P.M., Bibby K., Haramoto E., Hewitt J., Huygens F., Gyawali P., Korajkic A., Riddel S., Sherchan S.P., Simpson S.L., Sirikanchana K., Symonds E.M., Verhagen R., Vasan S.S., Kitajima M., Bivins A. Decay of SARS-CoV-2 and surrogate murine hepatitis virus RNA in untreated wastewater to inform application in wastewater-based epidemiology. Environ. Res. 2020;191 doi: 10.1016/j.envres.2020.110092. - DOI - PMC - PubMed
-
- Ahmed W., Bivins A., Bertsch P.M., Bibby K., Gyawali P., Sherchan S.P., Simpson S.L., Thomas K.V., Verhagen R., Kitajima M., Mueller J.F., Korajkic A. Intraday variability of indicator and pathogenic viruses in 1-h and 24-h composite wastewater samples: implications for wastewater-based epidemiology. Environ. Res. 2021;193 doi: 10.1016/j.envres.2020.110531. - DOI - PMC - PubMed
-
- Betancourt W.Q., Schmitz B.W., Innes G.K., Prasek S.M., Pogreba Brown K.M., Stark E.R., Foster A.R., Sprissler R.S., Harris D.T., Sherchan S.P., Gerba C.P., Pepper I.L. COVID-19 containment on a college campus via wastewater-based epidemiology, targeted clinical testing and an intervention. Sci. Total Environ. 2021;779 doi: 10.1016/J.SCITOTENV.2021.146408. - DOI - PMC - PubMed
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

