DNMT3b protects centromere integrity by restricting R-loop-mediated DNA damage
- PMID: 35688824
- PMCID: PMC9187704
- DOI: 10.1038/s41419-022-04989-1
DNMT3b protects centromere integrity by restricting R-loop-mediated DNA damage
Abstract
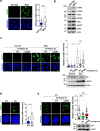
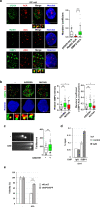
This study used DNA methyltransferase 3b (DNMT3b) knockout cells and the functional loss of DNMT3b mutation in immunodeficiency-centromeric instability-facial anomalies syndrome (ICF) cells to understand how DNMT3b dysfunction causes genome instability. We demonstrated that R-loops contribute to DNA damages in DNMT3b knockout and ICF cells. More prominent DNA damage signal in DNMT3b knockout cells was due to the loss of DNMT3b expression and the acquirement of p53 mutation. Genome-wide ChIP-sequencing mapped DNA damage sites at satellite repetitive DNA sequences including (peri-)centromere regions. However, the steady-state levels of (peri-)centromeric R-loops were reduced in DNMT3b knockout and ICF cells. Our analysis indicates that XPG and XPF endonucleases-mediated cleavages remove (peri-)centromeric R-loops to generate DNA beaks, causing chromosome instability. DNMT3b dysfunctions clearly increase R-loops susceptibility to the cleavage process. Finally, we showed that DNA double-strand breaks (DSBs) in centromere are probably repaired by error-prone end-joining pathway in ICF cells. Thus, DNMT3 dysfunctions undermine the integrity of centromere by R-loop-mediated DNA damages and repair.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures






References
-
- Okano M, Bell DW, Haber DA, Li E. DNA methyltransferases Dnmt3a and Dnmt3b are essential for de novo methylation and mammalian development. Cell. 1999;99:247–57. - PubMed
-
- Li E, Bestor TH, Jaenisch R. Targeted mutation of the DNA methyltransferase gene results in embryonic lethality. Cell. 1992;69:915–26. - PubMed
-
- Dodge JE, Okano M, Dick F, Tsujimoto N, Chen T, Wang S, et al. Inactivation of Dnmt3b in mouse embryonic fibroblasts results in DNA hypomethylation, chromosomal instability, and spontaneous immortalization. J Biol Chem. 2005;280:17986–91. - PubMed
-
- Xu GL, Bestor TH, Bourc’his D, Hsieh CL, Tommerup N, Bugge M, et al. Chromosome instability and immunodeficiency syndrome caused by mutations in a DNA methyltransferase gene. Nature. 1999;402:187–91. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

