Interpretation of gut microbiota data in the 'eye of the beholder': A commentary and re-evaluation of data from 'Impacts of radiation exposure on the bacterial and fungal microbiome of small mammals in the Chernobyl Exclusion Zone'
- PMID: 35694772
- PMCID: PMC9541917
- DOI: 10.1111/1365-2656.13667
Interpretation of gut microbiota data in the 'eye of the beholder': A commentary and re-evaluation of data from 'Impacts of radiation exposure on the bacterial and fungal microbiome of small mammals in the Chernobyl Exclusion Zone'
Abstract
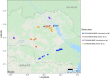
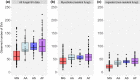
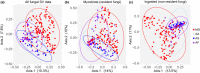
Evidence that exposure to environmental pollutants can alter the gut microbiota composition of wildlife includes studies of rodents exposed to radionuclides. Antwis et al. (2021) used amplicon sequencing to characterise the gut microbiota of four species of rodent (Myodes glareolus, Apodemus agrarius, A. flavicollis and A. sylvaticus) inhabiting the Chernobyl Exclusion Zone (CEZ) to examine possible changes in gut bacteria (microbiota) and gut fungi (mycobiota) associated with exposure to radionuclides and whether the sample type (from caecum or faeces) affected the analysis. The conclusions derived from the analyses of gut mycobiota are based on data that represent a mixture of ingested fungi (e.g. edible macrofungi, polypores, lichens and ectomycorrhizae) and gut mycobiota (e.g. microfungi and yeasts), which mask the patterns of inter- and intraspecific variation in the authentic gut mycobiota. Implying that 'faecal samples are not an accurate indicator of gut composition' creates an unnecessary controversy about faecal sampling because the comparison of samples from the caecum and faeces confounds many other possible drivers (including different animals from different locations, sampled in different years) of variation in gut microbiota. It is relevant also that Antwis et al.'s (2021) data lack statistical power to detect an effect of exposure to radionuclides on the gut microbiota because (1) all of their samples of Apodemus mice had experienced a medium or high total absorbed dose rate and (2) they did not collect samples of bank voles (M. glareolus) from replicate contaminated and uncontaminated locations. Discussion of Antwis et al.'s (2021) analysis, especially the claims presented in the Abstract, is important to prevent controversy about the outcome of research on the biological impacts of wildlife inhabiting the CEZ.
Keywords: amplicon sequencing; diet; microbiota; mycobiota; radiation effects.
© 2022 The Authors. Journal of Animal Ecology published by John Wiley & Sons Ltd on behalf of British Ecological Society.
Conflict of interest statement
The authors declare no conflict of interest.
Figures

Similar articles
-
Impacts of radiation exposure on the bacterial and fungal microbiome of small mammals in the Chernobyl Exclusion Zone.J Anim Ecol. 2021 Sep;90(9):2172-2187. doi: 10.1111/1365-2656.13507. Epub 2021 May 21. J Anim Ecol. 2021. PMID: 33901301
-
Skin and gut microbiomes of a wild mammal respond to different environmental cues.Microbiome. 2018 Nov 26;6(1):209. doi: 10.1186/s40168-018-0595-0. Microbiome. 2018. PMID: 30477569 Free PMC article.
-
Comparable response of wild rodent gut microbiome to anthropogenic habitat contamination.Mol Ecol. 2021 Jul;30(14):3485-3499. doi: 10.1111/mec.15945. Epub 2021 May 27. Mol Ecol. 2021. PMID: 33955637
-
The Role of Early Life Gut Mycobiome on Child Health.Adv Nutr. 2024 Mar;15(3):100185. doi: 10.1016/j.advnut.2024.100185. Epub 2024 Feb 3. Adv Nutr. 2024. PMID: 38311313 Free PMC article. Review.
-
Fungi in the healthy human gastrointestinal tract.Virulence. 2017 Apr 3;8(3):352-358. doi: 10.1080/21505594.2016.1247140. Epub 2016 Oct 13. Virulence. 2017. PMID: 27736307 Free PMC article. Review.
Cited by
-
Seasonal shifts and land-use impact: unveiling the gut microbiomes of bank voles (Myodes glareolus) and common voles (Microtus arvalis).FEMS Microbiol Ecol. 2024 Nov 23;100(12):fiae159. doi: 10.1093/femsec/fiae159. FEMS Microbiol Ecol. 2024. PMID: 39611357 Free PMC article.
References
-
- Abt, K. F. , & Bock, W. F. (1998). Seasonal variations of diet composition in farmland field mice Apodemus spp. and bank voles Clethrionomys glareolus . Acta Theriologica, 43, 379–389. 10.4098/AT.arch.98-49 - DOI
-
- Albenberg, L. , Esipova, T. V. , Judge, C. P. , Bittinger, K. , Chen, J. , Laughlin, A. , Grunberg, S. , Baldassano, R. N. , Lewis, J. D. , Li, H. , Thom, S. R. , Bushman, F. D. , Vinogradov, S. A. , & Wu, G. D. (2014). Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota in humans and mice. Gastroenterology, 147, 1055–1063. 10.1053/j.gastro.2014.07.020 - DOI - PMC - PubMed
-
- Antwis, R. E. , Beresford, N. A. , Jackson, J. A. , Fawkes, R. , Barnett, C. L. , Potter, E. , Walker, L. , Gaschak, S. , & Wood, M. D. (2021). Impacts of radiation on the bacterial and fungal microbiome of small mammals in the Chernobyl Exclusion Zone. Journal of Animal Ecology, 9, 2172–2187. 10.1111/1365-2656.13507 - DOI - PubMed
-
- Barelli, C. , Albanese, D. , Stumpf, R. M. , Asangba, A. , Donati, C. , Rovero, F. , & Hauffe, H. C. (2020). The gut microbiota communities of wild arboreal and ground‐feeding tropical primates are affected differently by habitat disturbance. MSystems, 5, 1–18. 10.1128/msystems.00061-20 - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

