Accelerated Identification of Cell Active KRAS Inhibitory Macrocyclic Peptides using Mixture Libraries and Automated Ligand Identification System (ALIS) Technology
- PMID: 35707970
- PMCID: PMC9289880
- DOI: 10.1021/acs.jmedchem.2c00154
Accelerated Identification of Cell Active KRAS Inhibitory Macrocyclic Peptides using Mixture Libraries and Automated Ligand Identification System (ALIS) Technology
Abstract
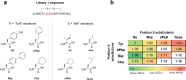
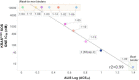
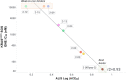
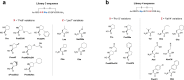
Macrocyclic peptides can disrupt previously intractable protein-protein interactions (PPIs) relevant to oncology targets such as KRAS. Early hits often lack cellular activity and require meticulous improvement of affinity, permeability, and metabolic stability to become viable leads. We have validated the use of the Automated Ligand Identification System (ALIS) to screen oncogenic KRASG12D (GDP) against mass-encoded mini-libraries of macrocyclic peptides and accelerate our structure-activity relationship (SAR) exploration. These mixture libraries were generated by premixing various unnatural amino acids without the need for the laborious purification of individual peptides. The affinity ranking of the peptide sequences provided SAR-rich data sets that led to the selection of novel potency-enhancing substitutions in our subsequent designs. Additional stability and permeability optimization resulted in the identification of peptide 7 that inhibited pERK activity in a pancreatic cancer cell line. More broadly, this methodology offers an efficient alternative to accelerate the fastidious hit-to-lead optimization of PPI peptide inhibitors.
Conflict of interest statement
The authors declare no competing financial interest.
Figures



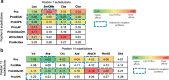

References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Chemical Information
Miscellaneous

