Programming a DNA tetrahedral nanomachine as an integrative tool for intracellular microRNA biosensing and stimulus-unlocked target regulation
- PMID: 35711289
- PMCID: PMC9194454
- DOI: 10.1016/j.mtbio.2022.100276
Programming a DNA tetrahedral nanomachine as an integrative tool for intracellular microRNA biosensing and stimulus-unlocked target regulation
Abstract
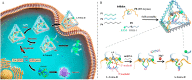
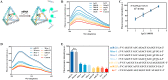
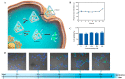
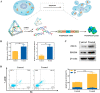
The synchronous detection and regulation of microRNAs (miRNAs) are essential for the early tumor diagnosis and treatment but remains a challenge. An integrative DNA tetrahedral nanomachine was self-assembled for sensitive detection and negative feedback regulation of intracellular miRNAs. This nanomachine comprised a DNA tetrahedron nanostructure as the framework, and a miRNA inhibitor-controlled allosteric DNAzyme as the core. The DNA tetrahedron brought the DNAzyme and the substrate in spatial proximity and facilitated the cellular uptake of DNAzyme. In allosteric regulation of DNAzyme, the locked tetrahedral DNAzyme (L-tetra-D) and active tetrahedral DNAzyme (A-Tetra-D) were controlled by miRNA inhibitor. The combination of miRNA inhibitor and target could trigger the conformational change from L-tetra-D to A-Tetra-D. A-Tetra-D cleaved the substrate and released fluorescence for intracellular miRNA biosensing. The DNA tetrahedral nanomachine showed excellent sensitivity (with detection limit down to 0.77 pM), specificity (with one-base mismatch discrimination), biocompatibility and stability. Simultaneously, miRNA stimulus-unlocked inhibitor introduced by our nanomachine exhibited the synchronous regulation of target cells, of which regulatory performance has been verified by the upregulated levels of downstream genes/proteins and the increased cellular apoptosis. Our study demonstrated that the DNA tetrahedral nanomachine is a promising biosense-and-treat tool for the synchronous detection and regulation of intracellular miRNA, and is expected to be applied in the early diagnosis and tailored management of cancers.
Keywords: DNA nanomaterials; DNA tetrahedron; DNAzyme; Target regulation; microRNA detection.
© 2022 Published by Elsevier Ltd.
Conflict of interest statement
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



References
-
- Lu D., Thum T. RNA-based diagnostic and therapeutic strategies for cardiovascular disease. Nat. Rev. Cardiol. 2019;16(11):661–674. - PubMed
-
- Rupaimoole R., Slack F.J. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat. Rev. Drug Discov. 2017;16(3):203–222. - PubMed
-
- Nucera S., Giustacchini A., Boccalatte F., Calabria A., Fanciullo C., Plati T., Ranghetti A., Garcia-Manteiga J., Cittaro D., Benedicenti F., Lechman E.R., Dick J.E., Ponzoni M., Ciceri F., Montini E., Gentner B., Naldini L. miRNA-126 orchestrates an oncogenic program in B cell precursor acute lymphoblastic leukemia. Cancer Cell. 2016;29(6):905–921. - PubMed
-
- Pichiorri F., Suh S.S., Rocci A., De Luca L., Taccioli C., Santhanam R., Zhou W., Benson D.J., Hofmainster C., Alder H., Garofalo M., Di Leva G., Volinia S., Lin H.J., Perrotti D., Kuehl M., Aqeilan R.I., Palumbo A., Croce C.M. Downregulation of p53-inducible microRNAs 192, 194, and 215 impairs the p53/MDM2 autoregulatory loop in multiple myeloma development. Cancer Cell. 2016;30(2):349–351. - PubMed
LinkOut - more resources
Full Text Sources

