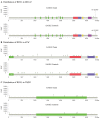
Rare germline deleterious variants increase susceptibility for lung cancer
- PMID: 35717579
- PMCID: PMC9558843
- DOI: 10.1093/hmg/ddac123
Rare germline deleterious variants increase susceptibility for lung cancer
Abstract
Although multiple common susceptibility loci for lung cancer (LC) have been identified by genome-wide association studies, they can explain only a small portion of heritability. The etiological contribution of rare deleterious variants (RDVs) to LC risk is not fully characterized and may account for part of the missing heritability. Here, we sequenced the whole exomes of 2777 participants from the Environment and Genetics in Lung cancer Etiology study, a homogenous population including 1461 LC cases and 1316 controls. In single-variant analyses, we identified a new RDV, rs77187983 [EHBP1, odds ratio (OR) = 3.13, 95% confidence interval (CI) = 1.34-7.30, P = 0.008] and replicated two previously reported RDVs, rs11571833 (BRCA2, OR = 2.18; 95% CI = 1.25-3.81, P = 0.006) and rs752672077 (MPZL2, OR = 3.70, 95% CI = 1.04-13.15, P = 0.044). In gene-based analyses, we confirmed BRCA2 (P = 0.007) and ATM (P = 0.014) associations with LC risk and identified TRIB3 (P = 0.009), involved in maintaining genome stability and DNA repair, as a new candidate susceptibility gene. Furthermore, cases were enriched with RDVs in homologous recombination repair [carrier frequency (CF) = 22.9% versus 19.5%, P = 0.017] and Fanconi anemia (CF = 12.5% versus 10.2%, P = 0.036) pathways. Our results were not significant after multiple testing corrections but were enriched in cases versus controls from large scale public biobank resources, including The Cancer Genome Atlas, FinnGen and UK Biobank. Our study identifies novel candidate genes and highlights the importance of RDVs in DNA repair-related genes for LC susceptibility. These findings improve our understanding of LC heritability and may contribute to the development of risk stratification and prevention strategies.
Published by Oxford University Press 2022.
Figures
References
-
- Siegel, R.L., Miller, K.D., Fuchs, H.E. and Jemal, A. (2021) Cancer statistics, 2021. CA Cancer J. Clin., 71, 7–33. - PubMed
-
- Islami, F., Goding Sauer, A., Miller, K.D., Siegel, R.L., Fedewa, S.A., Jacobs, E.J., McCullough, M.L., Patel, A.V., Ma, J., Soerjomataram, I. et al. (2018) Proportion and number of cancer cases and deaths attributable to potentially modifiable risk factors in the United States. CA Cancer J. Clin., 68, 31–54. - PubMed
-
- McKay, J.D., Hung, R.J., Han, Y., Zong, X., Carreras-Torres, R., Christiani, D.C., Caporaso, N.E., Johansson, M., Xiao, X., Li, Y. et al. (2017) Large-scale association analysis identifies new lung cancer susceptibility loci and heterogeneity in genetic susceptibility across histological subtypes. Nat. Genet., 49, 1126–1132. - PMC - PubMed
Publication types
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous