Capsaicin regulates lipid metabolism through modulation of bile acid/gut microbiota metabolism in high-fat-fed SD rats
- PMID: 35721805
- PMCID: PMC9180124
- DOI: 10.29219/fnr.v66.8289
Capsaicin regulates lipid metabolism through modulation of bile acid/gut microbiota metabolism in high-fat-fed SD rats
Abstract
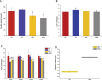
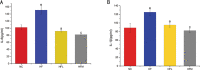
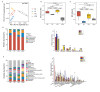
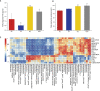
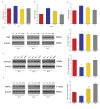
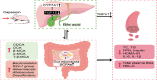
Capsaicin (CAP) is one of the active ingredients found in chili peppers and has been shown to reduce fat. This study aimed to explore the mechanisms of CAP activity by investigating intestinal microorganisms and bile acids (BAs). This study utilized 16S RNA sequencing to detect gut microbiota in cecal contents, and BAs in Sprague Dawley (SD) rats were also investigated. The results showed that 1) CAP increased the levels of chenodeoxycholic acid (CDCA), deoxycholic acid (DCA), β-muricholic acid (β-MCA), and tauro-β-muricholic acid sodium salt (T-β-MCA), which can regulate farnesoid X receptor (FXR) to inhibit Fgf15, increased CYP7A1 expression to lower triglycerides (TG) and total cholesterol (TC); 2) CAP decreased the abundance of Firmicutes and promoted the presence of specific fermentative bacterial populations, like Akkermansia; meanwhile, less optimal dose can reduce Desulfovibrio; 3) CAP decreased inflammatory factors IL-6 and IL-1β, and increased transient receptor potential channel of vanilloid subtype 1 (TRPV1) to regulate lipid metabolism, fasting plasma glucose and insulin resistance. In conclusion, CAP can reduce fat accumulation by regulating BAs, microorganisms, and short-chain fatty acids.
Keywords: bile acid; capsaicin; gut microbiota; high-fat diet; metabolism.
© 2022 Ting Gong et al.
Conflict of interest statement
All authors declare that they have no conflict of interest. The authors have not received any funding or benefits from industry or elsewhere to conduct this study
Figures
References
LinkOut - more resources
Full Text Sources
Miscellaneous

