N6-methyladenosine modification of the Aedes aegypti transcriptome and its alteration upon dengue virus infection in Aag2 cell line
- PMID: 35725909
- PMCID: PMC9209429
- DOI: 10.1038/s42003-022-03566-8
N6-methyladenosine modification of the Aedes aegypti transcriptome and its alteration upon dengue virus infection in Aag2 cell line
Abstract
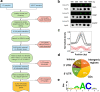
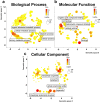
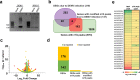
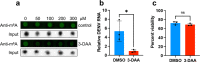
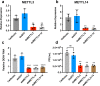
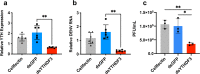
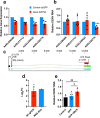
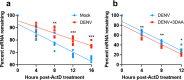
The N6-methyladenosine (m6A) modification of RNA has been reported to affect viral infections. Studies have confirmed the role of m6A in replication of several vector-borne flaviviruses, including dengue virus (DENV), in mammalian cells. Here, we explored the role of m6A in DENV replication in the mosquito Aedes aegypti Aag2 cell line. We first determined the presence of m6A on the RNAs from mosquito cells and using methylated RNA immunoprecipitation and sequencing (MeRIP-Seq) identified m6A modification of the mosquito transcriptome and those that changed upon DENV infection. Depletion of m6A methyltransferases and the m6A binding protein YTHDF3 RNAs decreased the replication of DENV. In particular, we found that the Ae. aegypti ubiquitin carrier protein 9 (Ubc9) is m6A modified and its expression increases after DENV infection. Silencing of the gene and ectopic expression of Ubc9 led to reduced and increased DENV replication, respectively. The abundance of Ubc9 mRNA and its stability were reduced with the inhibition of m6A modification, implying that m6A modification of Ubc9 might enhance expression of the gene. We also show that the genome of DENV is m6A modified at five sites in mosquito cells. Altogether, this work reveals the involvement of m6A modification in Ae. aegypti-DENV interaction.
© 2022. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

Similar articles
-
Transcriptome analysis of Aedes aegypti Aag2 cells in response to dengue virus-2 infection.Parasit Vectors. 2020 Aug 17;13(1):421. doi: 10.1186/s13071-020-04294-w. Parasit Vectors. 2020. PMID: 32807211 Free PMC article.
-
Role of Vigilin and RACK1 in dengue virus-Aedes aegypti-Wolbachia interactions.mSphere. 2025 Jan 28;10(1):e0048224. doi: 10.1128/msphere.00482-24. Epub 2024 Dec 23. mSphere. 2025. PMID: 39714171 Free PMC article.
-
Interaction of the Wolbachia surface protein with a novel pro-viral protein from Aedes aegypti.mBio. 2025 Jan 8;16(1):e0148624. doi: 10.1128/mbio.01486-24. Epub 2024 Nov 22. mBio. 2025. PMID: 39576110 Free PMC article.
-
The Involvement of Atlastin in Dengue Virus and Wolbachia Infection in Aedes aegypti and Its Regulation by aae-miR-989.Microbiol Spectr. 2022 Oct 26;10(5):e0225822. doi: 10.1128/spectrum.02258-22. Epub 2022 Sep 27. Microbiol Spectr. 2022. PMID: 36165808 Free PMC article.
-
Decoding epitranscriptomic regulation of viral infection: mapping of RNA N6-methyladenosine by advanced sequencing technologies.Cell Mol Biol Lett. 2024 Mar 27;29(1):42. doi: 10.1186/s11658-024-00564-y. Cell Mol Biol Lett. 2024. PMID: 38539075 Free PMC article. Review.
Cited by
-
Dynamic transcriptome analyses reveal m6A regulated immune non-coding RNAs during dengue disease progression.Heliyon. 2023 Jan 4;9(1):e12690. doi: 10.1016/j.heliyon.2022.e12690. eCollection 2023 Jan. Heliyon. 2023. PMID: 36685392 Free PMC article.
-
Towards a Universal Translator: Decoding the PTMs That Regulate Orthoflavivirus Infection.Viruses. 2025 Feb 19;17(2):287. doi: 10.3390/v17020287. Viruses. 2025. PMID: 40007042 Free PMC article. Review.
-
N6-methyladenosine RNA modification promotes Severe Fever with Thrombocytopenia Syndrome Virus infection.PLoS Pathog. 2024 Nov 25;20(11):e1012725. doi: 10.1371/journal.ppat.1012725. eCollection 2024 Nov. PLoS Pathog. 2024. PMID: 39585899 Free PMC article.
-
N-acetyltransferase 10 regulates alphavirus replication via N4-acetylcytidine (ac4C) modification of the lymphocyte antigen six family member E (LY6E) mRNA.J Virol. 2024 Jan 23;98(1):e0135023. doi: 10.1128/jvi.01350-23. Epub 2024 Jan 3. J Virol. 2024. PMID: 38169284 Free PMC article.
-
Differing Transcriptomic Responses in High Titer versus Low Titer Aedes aegypti Mosquitoes after Oral Infection with Sindbis Virus.Viruses. 2024 Sep 19;16(9):1487. doi: 10.3390/v16091487. Viruses. 2024. PMID: 39339963 Free PMC article.
References
-
- Park CH, Hong K. Epitranscriptome: m6A and its function in stem cell biology. Genes Genomics. 2017;39:371–378. doi: 10.1007/s13258-016-0507-2. - DOI
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

