Sequence learning, prediction, and replay in networks of spiking neurons
- PMID: 35727857
- PMCID: PMC9273101
- DOI: 10.1371/journal.pcbi.1010233
Sequence learning, prediction, and replay in networks of spiking neurons
Abstract
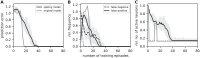
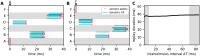
Sequence learning, prediction and replay have been proposed to constitute the universal computations performed by the neocortex. The Hierarchical Temporal Memory (HTM) algorithm realizes these forms of computation. It learns sequences in an unsupervised and continuous manner using local learning rules, permits a context specific prediction of future sequence elements, and generates mismatch signals in case the predictions are not met. While the HTM algorithm accounts for a number of biological features such as topographic receptive fields, nonlinear dendritic processing, and sparse connectivity, it is based on abstract discrete-time neuron and synapse dynamics, as well as on plasticity mechanisms that can only partly be related to known biological mechanisms. Here, we devise a continuous-time implementation of the temporal-memory (TM) component of the HTM algorithm, which is based on a recurrent network of spiking neurons with biophysically interpretable variables and parameters. The model learns high-order sequences by means of a structural Hebbian synaptic plasticity mechanism supplemented with a rate-based homeostatic control. In combination with nonlinear dendritic input integration and local inhibitory feedback, this type of plasticity leads to the dynamic self-organization of narrow sequence-specific subnetworks. These subnetworks provide the substrate for a faithful propagation of sparse, synchronous activity, and, thereby, for a robust, context specific prediction of future sequence elements as well as for the autonomous replay of previously learned sequences. By strengthening the link to biology, our implementation facilitates the evaluation of the TM hypothesis based on experimentally accessible quantities. The continuous-time implementation of the TM algorithm permits, in particular, an investigation of the role of sequence timing for sequence learning, prediction and replay. We demonstrate this aspect by studying the effect of the sequence speed on the sequence learning performance and on the speed of autonomous sequence replay.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures









References
-
- Lashley KS. The problem of serial order in behavior. vol. 21. Bobbs-Merrill; 1951.
-
- Hawkins J, Blakeslee S. On intelligence: How a new understanding of the brain will lead to the creation of truly intelligent machines. Macmillan; 2007.
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Miscellaneous

