Dysregulated Gene Expression in Lymphoblasts from Parkinson's Disease
- PMID: 35736800
- PMCID: PMC9230639
- DOI: 10.3390/proteomes10020020
Dysregulated Gene Expression in Lymphoblasts from Parkinson's Disease
Abstract
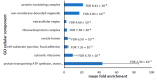
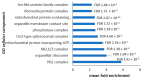
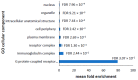
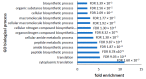
Parkinson's disease is the second largest neurodegenerative disease worldwide and is caused by a combination of genetics and environment. It is characterized by the death of neurons in the substantia nigra of the brain but is not solely a disease of the brain, as it affects multiple tissues and organs. Studying Parkinson's disease in accessible tissues such as skin and blood has increased our understanding of the disease's pathogenesis. Here, we used lymphoblast cell lines generated from Parkinson's disease patient and healthy age- and sex-matched control groups and obtained their whole-cell transcriptomes and proteomes. Our analysis revealed, in both the transcriptomes and the proteomes of PD cells, a global downregulation of genes involved in protein synthesis, as well as the upregulation of immune processes and sphingolipid metabolism. In contrast, we discovered an uncoupling of mRNA and protein expression in processes associated with mitochondrial respiration in the form of a general downregulation in associated transcripts and an upregulation in proteins. Complex V was different to the other oxidative phosphorylation complexes in that the levels of its associated transcripts were also lower, but the levels of their encoded polypeptides were not elevated. This may suggest that further layers of regulation specific to Complex V are in play.
Keywords: Parkinson’s disease; cell models; mitochondria; oxidative phosphorylation; protein synthesis; proteome; transcriptome.
Conflict of interest statement
The authors declare no conflict of interest.
Figures



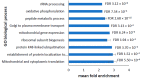
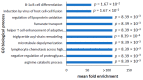

References
-
- Parkinson J. An Essay on the Shaking Palsy. Sherwood, Neely, and Jones; London, UK: 1817.
Grants and funding
LinkOut - more resources
Full Text Sources

